Академический Документы
Профессиональный Документы
Культура Документы
James Lee Enzyme Kinetics Solution
Загружено:
VPrasarnth RaajИсходное описание:
Авторское право
Доступные форматы
Поделиться этим документом
Поделиться или встроить документ
Этот документ был вам полезен?
Это неприемлемый материал?
Пожаловаться на этот документАвторское право:
Доступные форматы
James Lee Enzyme Kinetics Solution
Загружено:
VPrasarnth RaajАвторское право:
Доступные форматы
BK10110302 V.
PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.1
Total volume = 44 + 5 + 1= 50ml
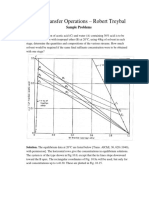
From the graph the equation obtained
y=0.033 + 0.03a 1
m= 0.033 mol / ml . min
a) Activity of the - glucosidase
0.033 x 50= 1.65 mumol / min
i)
= 1.65mumol/min
0.1 mg/ml x 0.1ml
= 165 units/mg protein
ii)
= 1.65 mumol/min
1ml of enzyme
= 1.65 units/ml of enzyme
b) Initial rate of reaction 0.033 mumol/ mL.min
S vs t Graph
1.2
y = 0.033x + 0.0391
1
0.8
0.6
0.4
0.2
0
0
10
15
20
25
30
35
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.2
a) Michaelis-Menten approach
The rate of product formation.
d[p]
dt
Since the enzyme is preferred,
Make E as the subject,
Since forward reaction = backward reaction.
( )
ubstitute
into
:
[(
(
Make
) (
)( )
)] ( )
)( )
)( )
as a subject:
(
)( )
)( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
(
)( )
)( )
[(
(
(
(
ub
into
)( )
)( )
)( )
)( )( )
)( )( )
as a subject,
(
)(
( )( ))
)
(
into
)( )( )
)( )( )
)(
)
(
ub
)( )
)
Make
)( )]
( )( ))
)( )( )
( )
( )( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
d[p]
dt
Since
)( )( )
)
( )
( )( )
)( )( )
)
( )
( )( )
)
( )( )
)
( )
( )( )
b) Since [ ]
[ ]
d[p]
dt
( )( )
( )( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.3
(a) E+S k1
(ES)1
(ES)1 k3 (ES)2
(ES)2 k2 E+P
V=
[ ]
= k5 [ES] 2
[E0] = [E] + [ES] + [ES]2
[E] = [E0]-[ES]-[ES]2
k2 = [E] [S]
k1 [ES]1
k2 [ES]1 = [Eo] [S] [ES]1 [S]
k1
[ES]1 ( k2 + [S] ) = [E0] [S] [ES]2 [S]
k1
[ES]1 = [E0] [S] [ES]2 [S]
k2 + [S]
k1
k4 = [ES]1
k3
[ES]2
k4 [ES]2 = [E0] [S] [ES]2 [S]
k3
k2 + [S]
k1
[ES]2 ( k2 k4 + k4 [S] ) = [E0] [S] [ES]2 [S]
k1k3
k3
[ES]2 ( k2 k4 + k4 [S] + [S] )= [E0] [S]
k1 k3
k3
[ES]2 = [E0] [S]
k2 k4 + k4 [S] + [S]
k1 k3
k3
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
V= d [P] = k5 [E0] [S]
dt
k2 k4 + k4 [S] + [S]
k1 k3
k3
=
Vm [S]
k2 k4 + k4 [S] + [S]
k1 k3
k3
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.4
a) Michaelis-Menten approach
The rate of product formation.
d[p]
dt
Since the enzyme is preferred,
Make E as the subject,
Since forward reaction = backward reaction.
( )
ubstitute
into
:
[(
Make
) (
)( )
)] ( )
)( )
)( )
as a subject:
(
)( )
)( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
(
)( )
)( )
[(
(
(
(
ub
into
)( )
)( )
)( )
)( )( )
)( )( )
as a subject,
(
)(
( )( ))
)
(
into
)( )( )
)( )( )
)(
)
(
ub
)( )
)
Make
)( )]
( )( ))
)( )( )
( )
( )( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
d[p]
dt
Since
)( )( )
)
( )
( )( )
)( )( )
)
( )
( )( )
)
( )( )
)
( )
( )( )
b) Briggs-Haldane approach
The rate of product formation,
d(p)
dt
Since the enzyme is preferred,
Make as a subject,
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Substrate consumption,
d(
( )( )
dt
d(
dt
ubstitute
into
)(
)(
)(
)( )
( )
( )
( )
( )
ubstitute
)( )
( )( )
)( )
( )(
( ))
( )
( )
( )(
( )
( )(
( )(
(
( ))
into
)( )
:
(
)(
)(
)(
( )
)( )
( )(
(
( ) ( )
( )
( ) ( )(
(
( )
( ))
)
( ))
( )
)( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
)(
)(
( )
( ) ( )
( ) ( )
(
ubstitute
)(
( ))
( ) ( )(
( )
)(
( )
( )
( ))
( ) ( )
( ) ( )
(
( ) ( )
( )
)(
( )
( ))
into
d(p)
dt
( ) ( )
(
( ) ( )
( )
)(
( )
( ))
v
d(p)
dt
v
(
( ) ( )
( ) ( )
( )
)(
( )
( ))
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.5
Lineweaver- Burk Plot
x-intercept= - 1
km
y-intercept= 1/ V more
Equation obtained y= 0.0172 x + 3.6342
y-intercept = 3.6342= 1/ V max
V max = 0.275
x-intercept , y= 0
0.0172x + 3.6342=0
0.0172x = -3.6342
x= -211.291
x= -1/km
km = 1/211.291
= 0.00473
Longmuir Plot
Equation obtained y= 3.3133x + 0.0191
1/Vm = m = 3.3133
Vm=0.302
y-intercept= km/Vm = 0.0191
Km = 0,0191x 0.302
= 0.00577
Eadie-Hofstee Plot
Equation obtained y= -0.0043x + 0.2645
-Km = m = -3.3133
Km=0.302
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
y-intercept= Vm = 0.2645
Non-Linear Regression Procedure
From the graph, Vm=0.2
Vmax = 0.1,
Km=0.0032
Data for Graph plot :
Langmuir Plot
s
s/v
0.0032 0.028829
0.0049 0.033108
0.0062 0.043357
0.008 0.048193
0.0095
0.0475
Lineweaver-Burk Plot
1/s
1/v
312.5
9.009009
204.0816 6.756757
161.2903 6.993007
125
6.024096
105.2632
5
Eadie-Hofstee Plot
v/s
v
34.6875
0.111
30.20408 0.148
23.06452 0.143
20.75
0.166
21.05263
0.2
Non-Linear Regression Plot
S
v
0.0032
0.111
0.0049
0.148
0.0062
0.143
0.008
0.166
0.0095
0.2
Kinetic Parameters
Vmax
Km
Langmuir
0.2750
0.0047
Lineweaver-Burk
0.0191
0.0057
Eadie-Hofstee
0.2645
0.0043
Non-Linear Regression 0.2000
0.0032
Type of Plot
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Langmuir Plot
0.06
y = 3.3133x + 0.0191
0.05
0.04
0.03
0.02
0.01
0
0
0.002
0.004
0.006
0.008
0.01
Lineweaver Burk Plot
10
9
8
7
6
5
4
3
2
1
0
y = 0.0172x + 3.6342
50
100
150
200
250
300
350
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Eadie-Hofstee Plot
0.25
0.2
0.15
y = -0.0043x + 0.2645
0.1
0.05
0
0
10
15
20
25
30
35
40
Non-Linear Regression Procedure
0.25
0.2
0.15
0.1
0.05
0
0
0.002
0.004
0.006
0.008
0.01
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.6
Rate of product formation
( )
Enzyme is preserved,
d( )
dt
(
Assumptions:
[ ]small,
neg igib e
( )( )
)(
)(
(
(
into
))( )
)
( )
(
(
)
)
( ) )
( )
( )
(
( )
d(p)
dt
(
(
( )
into
( )
(
( )( )
(
)
Substitute
v
( )( )
Substitute equation
(
()
( )
( )
)
)
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Dividing
[(
with the value of
( )
( )
]/
v s
( )
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.7
a) FCs0 - FCs + rSv = V
For Batch reactor F=0
rSv = V
[ ]
[ ]
[
]
[
= 60mol/m3.min
b) Equation obtained y = 6.3852x + 59.571
m = Vmax
= 6.3852
y- intercept = - Km = 59.571
Km = - 59.571
c) FCs0 - FCs + rSv = 0
FCs0 - FCs = - rSv = rpv
FCs0 - FCs =
F = 0.0001m3/min
V = 0.0003m3
( FCSo - FCs ) (Km + Cs)
= Vmax CsV
2
FCSo Km + FCSo Cs - FKm Cs FCs = Vmax CsV
(0.0001 (300)(200) + 0.0001(300)Cs 0.001(200)Cs 0.001Cs2 = 100 (0.0003)Cs )
6 + 0.03Cs 0.02Cs 0.001Cs2 = 0.03Cs
0.0001Cs2 + 0.02Cs 6 = 0
Cs=165mol/m3
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Data :
Cs t t/ln(Cso/Cs) (Cso-Cs)/ln(Cso/Cs)
1 1
0.175322
52.42135
5 5
1.221197
72.0506
10 10
2.940141
85.26409
20 20
7.385387
103.3954
Graph :
(Cso-Cs)/ln(Cso/Cs)
120
y = 6.3852x + 59.571
100
80
(Cso-Cs)/ln(Cso/Cs)
60
Linear ((CsoCs)/ln(Cso/Cs))
40
20
0
0
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.8
a) Km =0.01 mol/L
Cso = 3.4 x 10 -4 mol/L
Cs = 0.9 x 3.4 x 10-4
= 3.06 x 10-4 mol/L
t= 5minutes
[ ]
[ ]
( 3.4x 10-4 3.06 x 10-4) = Vmax (3.06 x 10-4)
S
0.01 + (3.06 x 10-4)
6.8 x 10-6 = Vmax ( 0.03)
Vmax = 2.27 x 10-4 mol/L-min
b) 6.8 x 10-6 x 15 = 1.02 x 10-4 mol/L
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.9
Km = 0.03mol/L
rmax = 13mol/L min 60 = 780mol/L hr
F=10L/Hr
Cs=10mol/L
F=10L/Hr
Cs=0.5mol/L
CSTR
a) V = ?
CSTR @ Stead State
FCs0 - FCs + rSv = 0
F (Cs0 - Cs ) = rpv
(
10 (10 0.5) =
V = 0.129 liter
b) Plug - Flow @ Stead State
Km ln
+ (Cs0 - Cs )
0.03 ln
+ (10 - 0.5 ) = 780t
= rmax t
9.95899 = 780t
t = 0.0123hr
t = V/F = 0.0123
V = 0.0123 10
= 0.123liter
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.10
Km = 10g/L
rmax = 7g/L.min
a)
F=0.5L/min
Cs0=50g/L
1L
F=0.5L/min
Cs1=?g/L
1L
F=0.5L/min
Cs2=? g/L
CSTR @ Steady State
FCs0 - FCs + rSv = 0
F (Cs0 - Cs ) = rpv
0.5 (50 Cs1) =
)
s
(1)
(25-0.5Cs1)(10+ Cs1)=7Cs1
250+25Cs1-5Cs1-0.5Cs12=7Cs1
0.5Cs12-13Cs1-250=0
Cs1=38.86g/L
0.5 (38.86 Cs2) =
)
s
(1)
(19.43-0.5 Cs2)(10+ Cs2)=7 Cs2
194.3+19.43Cs2-5Cs2-0.5Cs22=7Cs2
0.5Cs22-7.43Cs2-194.3 =0
Cs2=28.49g/L
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
b)
F=0.5L/min
Cs0=50g/L
2L
F=0.5L/min
Cs1=?g/L
CSTR @ Steady State
FCs0 - FCs + rSv = 0
F (Cs0 - Cs ) = rpv
0.5 (50 Cs1) =
)
s
(2)
(25-0.5Cs1)(10+ Cs1)=14Cs1
250+25Cs1-5Cs1-0.5Cs12=14Cs1
0.5Cs12-6Cs1-250=0
Cs1=29.15g/L
Since in the Cs in two reactor system is less than Cs in one reactor system, therefore two reactor
system is more efficient than one reactor system as it indicates more substrates have been
consumed to form products.
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.11
a) k1 [E] [S] = k2 [ES]
[ES] = k2
k1 [E] [S]
k1 [E] [S]
k2
k3 [E] [P] = k4 [EP]
[EP] = k3
k4 [E] [P]
k5 [ES][P] = k6 [ESP]
[ESP] = k5 [ES] [P]
k6
k7 [EP] [S] = k8 [ EPS ]
[EPS] = k7 [EP] [ S ]
K8
= k7 k3 [ S ]
k8 k4 [E] [P]
From,
[ESP] = k5 [P] k2
k6k1 [E] [S]
[E0] = [E] + [ES] + [EP] + [ESP] + [EPS]
[E0] = [ES] + [ESP] + [E] + [EP] + [EPS]
[E0] = [ES] + [ESP] + [E] + [EP] +
][ ]
[ ]
[E0] = [ES] + [ESP] + [E] + [EP] + (
[ ][ ]
[E0] = [ES] + [ESP] + [E] +
[ ]
[E0] = [ES] + [ESP] + [E] [
[E0] = [ES] +
[E0] = [ES] {1 +
][ ]
][
[ ]
[ ]
+(
[ ]
(
[ ]
[
[ ]
)]
[ ]
(
[ ]
)]}
)]
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
[ES] =
V=
[ ]
b)
Given:
[ ]
[ ]
KSP =
KSP =
[ESP] =
[ ]
[ ]
[ ]
)]
[ ]
)]
KPS =
[ ][ ][ ]
[
]
KPS =
[ ][ ][ ]
[
]
[ ][ ][ ]
[EPS] =
[ ][ ][ ]
][ ]
]
][ ]
]
[ESP] = [EPS]
KS KSP = KP KPS
=
[ ]
=
=
[
[ ]
[ ]
[ ]
]
[ ]
[ ]
[ ]
]
]
[ ]
[ ]
c)
Ks=Kps
Kp=Ksp
[ESP]=[EPS]
[ ]
(
( )
[
[ ]
[ ][ ]
)[
[ ]
][ ]
]
[ ]
[ ]
[ ]
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
[
][ ]
[ ])
[ ](
[ ]
[ ]
[ ]
[ ])
[ ]
[ ]
[ ])
(
(
Compare with
[ ]
[ ]
Hence, Vmax = (
Km=
d)
[ ])
[ ]
(
[ ])
[ ]
[ ]
[ ]
[ ]
*
[
[ ]
[ ]
[ ]
[ ]
[ ]
[ ]
[ ]
[ ]
[ ]
n(
[ ]
)
[ ]
[ ]
+
[ ] [ ]]
[ ] [ ]
[ ]
[ ]
[ ]
n(
[ ]
)
[ ]
n([ ] [ ])
(
[ ]
[ ]
)
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Y = mx+c
[ ]
)
]
n ([
Y=
M=
[
X= (
] [ ]
C=
So we can plot a graph of
[ ]
)
]
n ([
vs (
] [ ]
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.14
Rate:
rp = k9CES +k10CEIS1 +k10CEIS2
---- 1
Enzyme balance:
CEo = CE + CES
---- 2
CEo = CEIS1 + CES + CE
---- 3
CEo = CEIS2 + CEI + CE
---- 4
The equilibrium reaction equations are as follows:
CE Cs / CES = k2/k1
---- 5
CECI / CEI = K4/K3
---- 7
CESCI /CEIS1 = K6/K5
---- 6
CEICS / CEIS2 = K8/K7
---- 8
By rearranging Equation 5,
CE = (k2/k1) Cs CES
From Equation 2,
CEo = [(k2/k1)CE + 1] CES
CES = CEo /[( k2/k1)CS +1]
---- 9
By rearranging Equation 6,
CES = [(K6/K5)CI ] CEIS1
From Equation 3,
CEo = CEIS +CES + (k2/k1) Cs CES
= {CEIS1 + [1 + (k2/k1) Cs]( K6/K5)CI }CEIS1
= {1 + [1 + (K2/K1) Cs ]( K6/K5)CI } CEIS1
CEIS1 = CEo/ {1 + [1 + (k2/k1) Cs ]( K6/K5)CI }
By rearranging Equation 7,
CE = (K4/K3) CEI
By rearranging Equation 8,
CEI = K8/K7CS CEIS2
---- 10
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
From Equation 4,
CEo = CEIS2 + CEI + [(K4/K3)CI]CEI
= CEIS2 + [1 + (K4/K3)CI ]CEI
= CEIS2 + [1 + (K4/K3)CI ]( K8/K7)CS CEIS2
CEo = {1 + [1 + (K4/K3)CI ]( K8/K7)CS } CEIS2
CEIS2 = CEo / {1 + [1 + (K4/K3)CI ]( K8/K7)CS }
---- 11
From Equation 1, since rp = k9CES +k10CEIS1 +k10CEIS2,
By substituting Equation 9, 10 & 11 into Equation 1,
Therefore,
rp = k9 CEo /[( k2/k1)CS +1] + k10 CEo/ {1 + [1 + (k2/k1) Cs ]( K6/K5)CI } + k10 CEo / {1 + [1 + (K4/K3)CI
]( K8/K7)CS }
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.15
a) Based on the graphs
The y-intercept in Lineweaver Burk plot is almost the same.
Y-intercept => 3.8266; 3.6342
Whereas in Langmuir Plot
Two equations obtained
Y = 2.9883x + 0.0489
Y = 3.3133x + 0.0191
When y=0
X=
X=
X = -0.016
X = -0.005
In Line weaver Burk Plot and Langmuir Plot both indicates its a competitive inhibitor
Data :
Lineweaver
1/s
1/Vo
312.5
9.009009
204.0816 6.756757
161.2903 6.993007
125
6.024096
105.2632
5
Langmuir
s
s/Vo
0.0032 0.028829
0.0049 0.033108
0.0062 0.043357
0.008
0.048193
0.0095
0.0475
1/Vi
16.94915
14.08451
10.98901
9.009009
8
S/Vi
0.054237
0.069014
0.068132
0.072072
0.076
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Lineweaver-Burk Plot
20
18
16
14
12
10
8
6
4
2
0
y = 0.0439x + 3.8266
1/Vo
1/Vi
y = 0.0172x + 3.6342
Linear (1/Vo)
Linear (1/Vi)
100
200
300
400
Langmuir Plot
0.09
0.08
y = 2.9883x + 0.0489
0.07
0.06
s/Vo
y = 3.3133x + 0.0191
0.05
S/Vi
0.04
Linear (s/Vo)
0.03
Linear (S/Vi)
0.02
0.01
0
0
0.002
0.004
0.006
b) Y-intercept = 1/Vmax = 0.00489
Vmax = 1/0.00489 = 204.5 mol /L.min
Km/Vmax = 2.9883
Km=2.9883*204.5
=611mol/L
0.008
0.01
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.16
(a) E + S
ES + S
k5 E + P
(E0 ) = (E) +(ES) + (ESS)
(E) = (E0 ) (ES) (ESS) -------V=
=
(
( )
= k5 (ES) ------
)( )
)
) = (ES)(S)/ (k4 / k3)
K2 / k1 = (E)(S) / (ES)
K2/k1 (ES) = (E0)(S) (ES)(S)
(ES)((k2/k1) + (S)) = (E0)(S) (ES)(S)2 /
(ES)( (k2/k1) + (S)( ) ) =
(E0)(S) (ES)(S)2
(ES) ( (k2/k1) + (S)( ) + (S)2 ) =
(ES) =
(E0)(S)
(E0)(S) / (k2/k1) + (S)( ) + (S)2 --------
3
V=
=
( )
k5 (E0) (S) / (k2/k1) + (S)( ) + (S)2
Vm(S) / (k2/k1) + (S)( ) + (S)2
(b) At low substrate concentration,
1/ Vm = 3.1209
Vm = 0.3204
Km/Vm = 106.07
Km / 0.3204 = 106.7
K1m = 33.98
At high substrate concentration,
1/ Vm = 3.0574
Vm = 0.3271
1/ K1. Vm = 0.0032
1/ Km(3.0574) = 0.0032
Km = 102mol/L
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.17
V= 5L
Cso = 100 mmol/L
F = 1 L/hr
Cs = 10m mol/L
a) F (s0 FCs = rp V
1(100-10) = rp (5)
Rp = 18 m mol/ L.min
b) Find rp for each F and s
Equation obtained y= 0.0391x + 0.1641
M= 1/Vmax = 0.0391
Vmax= 25.57 m mol/L.min
Km/ Vm = 0.1641
Km= 4.197
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.18
[SO]1 = 0.1 mol/L
[S0]2 = 0.3mol/L
[ ][ ]
[
]
[ ][
]
]
[ ][
[ ][
]
]
[
[ ][
[
]
]
[ ][ ]
[
]
V1 =
= k5[ES1]
=3.5 [ES1] --V2 =
= k6[ES2]
=2.8 [ES2] ---
[E0] = [E] + [ES1] + [ES2]
[ ][
[E0] = [E] + [ES1] +
[E0] = [ES1] + [E] (1+
[E0] = [ES1] +
[E0] = [ES1] {1 +
[
[ES1] =
[
[
]
(1+
(1+
]
(
)}
[E0] = 0.05 mol/L
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Vmt = [S1]0 [S2] + K ln
[ ]
[ ]
3.5 [ES1] t = 0.1 [S1] +0.0714 ln [
3.5 [ES1] t + [S1] = 0.1 +0.0714 ln 0.1 - 0.0714 ln [S1]
3.5 [ES1] t + [S1] + 0.0644 = -0.0714 ln [S1]
ln[S1] =
[S1] =
---
[E0] = [E] + [ES1] + [ES2]
[E0] = [E] +
[ ][
[E0] = [E] (1+
[E0] =
[
]
[E0] = [E2] [
+ [ES2]
(1+
)+ [ES2]
[
(1+
Vmt = [S1]0 [S2] + KMln
)+ [ES2]
]
)+ 1]
]
2.8[ES1] t = 0.3 [S2] + 0.2207ln [
2.8[ES1] t + [S2] = 0.3 + 0.2207ln 0.3 0.2207ln [S2]
2.8[ES1] t + [S2] 0.0343 = 0.2207ln [S2]
ln[S2] =
[S2] = e
---
As [S1] increases, [ES1] also increases as in eq.3. [P1] also increases as in eq.1. This also occurs in
[S2]. As [S1] increases, [ES1] also increases as in eq.4. [P2] also increases as in eq.2
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.19
Data :
s
6.7
3.5
1.7
s/v
22.33333
14
10.625
Langmuir Plot
25
y = 2.3722x + 6.2429
20
15
10
5
0
0
1/Vm = 2.3722
Vmax = 0.4215 mumol/L.min
Km/Vm = 6.2429
Km = 6.2429(0.4215)
=2.63mumol/L
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Solution 2.20
a. Write the kinetic model.
Since the Michaelis constant KM is not affected by the presence of the inhibitor (which has shown on
the given table); then this enzyme reaction is noncompetitive inhibition reaction.
Kinetic Model:
k 1, k 2
E S
ES
k 3, k 4
E I
EI
k 5, k 6
EI S
EIS
k 7 ,k 8
ES I
ESI
k9
ES
EP
b. Derive the rate equation. State the assumptions.
Assumptions:
The dissociation constant for the first equilibrium reaction is the same as that of the third
equilibrium reaction.
The dissociation constant for the second equilibrium reaction is the same as that of the
fourth equilibrium reaction.
The two equilibrium reactions,
k
k2
K S 6 K IS
k1
k5
k
k4
K I 8 K SI
k3
k7
If the slower reaction, the product formation step, determines the rate of reaction according to
Michaelis-Menten assumption, the rate can be expressed as:
rP k 9 [ ES ]
(1)
The enzyme balance gives
[ E0 ] [ E ] [ ES ] [ EI ] [ ESI ]
(2)
k 9 [ ES ]
rP
[ E0 ] [ E ] [ ES ] [ EI ] [ ESI ]
(3)
Divide (1) by (2),
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Applied Law of mass action,
Ks
K 2 [ E ][S ]
[ E ][S ]
[ ES ]
K1
[ ES ]
KS
(4)
KI
K 4 [ E ][ I ]
[ E ][ I ]
[ EI ]
K3
[ EI ]
KI
(5)
KI
k8 [ ES ][ I ]
[ ES ][ I ]
[ ESI ]
k7
[ ESI ]
KI
(6)
Substitute (4), (5), (6) into (3),
[ E ][ S ]
KS
rP
[ E ][ S ] [ E ][ I ] [ ES ][ I ]
[ E0 ]
[E]
KS
KI
KI
k9
[ E ][ S ]
KS
rP
[ E ][ S ] [ E ][ I ] [ E ][ S ][ I ]
[ E0 ]
[E]
KS
KI
KS KI
k9
Eliminate [E],
[S ]
KS
rP
[ S ] [ I ] [ S ][ I ]
[ E0 ]k 9
1
KS KI KS KI
Substitute rPmax [ E0 ]k 9
rP
rPmax
[S ]
KS
[ S ] [ I ] [ S ][ I ]
1
KS KI KS KI
Multiply numerator and denominator by Ks,
rP
rPmax
[S ]
K [ I ] [ S ][ I ]
K S [S ] S
KI
KI
BK10110302 V.PRASARNTH RAAJ VEERA RAO BIOPROCESS
Rearranging,
rP
rPmax
[S ]
K [I ]
[ S ][ I ]
KS S
[S ]
KI
KI
rP
rPmax
[S ]
K S (1
[I ]
[I ]
) [ S ](1
)
KI
KI
Вам также может понравиться
- Sepa and PartechДокумент5 страницSepa and Partechdiana bunagan0% (1)
- Reviewer For Chemical Engineering Licensure Examination 3 Edition Solutions ManualДокумент56 страницReviewer For Chemical Engineering Licensure Examination 3 Edition Solutions ManualSherry Anne Ynciong Panganiban100% (4)
- Sample Problem #6Документ4 страницыSample Problem #6Dozdi100% (2)
- Astm A681 PDFДокумент14 страницAstm A681 PDFraulОценок пока нет
- British PetroleumДокумент10 страницBritish PetroleumVPrasarnth RaajОценок пока нет
- Extraction Problem Solving DrillДокумент2 страницыExtraction Problem Solving Drilledmark icalina40% (5)
- ScreeningДокумент30 страницScreeningPatricia de Leon100% (2)
- Astm F139Документ5 страницAstm F139diegomez84Оценок пока нет
- Balmes, Patricia R. - Enzyme KineticsДокумент11 страницBalmes, Patricia R. - Enzyme KineticsGlecie RasОценок пока нет
- Fire cable performance standardsДокумент3 страницыFire cable performance standardsRajan Varghese100% (1)
- Orca - Share - Media1575204849829 2Документ113 страницOrca - Share - Media1575204849829 2Lily Antonette AgustinОценок пока нет
- Kinetics (Gjjkkkgty)Документ5 страницKinetics (Gjjkkkgty)Chrysler Kane DepnagОценок пока нет
- Confined SpaceДокумент31 страницаConfined SpacegshdavidОценок пока нет
- Liquid Liquid ExtractionДокумент50 страницLiquid Liquid ExtractionArrianne Jaye Mata86% (14)
- Evaporator Heat and Mass Transfer CalculationsДокумент16 страницEvaporator Heat and Mass Transfer CalculationsArlene DeppОценок пока нет
- Day 2 200 ItemsДокумент25 страницDay 2 200 ItemsRobert DelfinОценок пока нет
- DocxДокумент2 страницыDocxDiana BunaganОценок пока нет
- BK10110302-Shuler Problems PDFДокумент13 страницBK10110302-Shuler Problems PDFVPrasarnth Raaj100% (2)
- Petroleum Assignment - UOP Q-Max Cumene Process (FULL)Документ436 страницPetroleum Assignment - UOP Q-Max Cumene Process (FULL)VPrasarnth Raaj100% (12)
- Problem Set - Enzymes From LehningerДокумент11 страницProblem Set - Enzymes From LehningervioletbrownОценок пока нет
- Chapter 8: Sterilization: Che 514A: Biochemical EngineeringДокумент9 страницChapter 8: Sterilization: Che 514A: Biochemical EngineeringEzekielОценок пока нет
- Safely Purging Air from Pipelines and VesselsДокумент7 страницSafely Purging Air from Pipelines and Vesselsgan_raoОценок пока нет
- مسائل وحلول ماسДокумент31 страницаمسائل وحلول ماسHmid Aljbre100% (4)
- Parker O-Ring Material Guide-2008Документ72 страницыParker O-Ring Material Guide-2008Ian Pillay50% (2)
- BIochem AssДокумент5 страницBIochem AssCheng PasionОценок пока нет
- Separation Process Compilation of Problem SetДокумент60 страницSeparation Process Compilation of Problem SetKaye Fabros100% (5)
- Test 1 Book Notes Examples-1 PDFДокумент65 страницTest 1 Book Notes Examples-1 PDFRalph Castino87% (15)
- ChE514A Cell Kinetics and Fermenter DesignДокумент116 страницChE514A Cell Kinetics and Fermenter DesignAnonymous 0zrCNQ55% (11)
- Soln KG Inert KG 5 - 1: Sample Problem #1Документ1 страницаSoln KG Inert KG 5 - 1: Sample Problem #1DozdiОценок пока нет
- ChE127 NUNEZ Assignment1Документ1 страницаChE127 NUNEZ Assignment1John Patrick Sanay NunezОценок пока нет
- Sample Problem #2Документ2 страницыSample Problem #2Dozdi67% (3)
- Stoichiometry of Microbial Growth and Product FormationДокумент10 страницStoichiometry of Microbial Growth and Product FormationfayeОценок пока нет
- Problems in Mass TransferДокумент3 страницыProblems in Mass TransferAngelica Joyce BenitoОценок пока нет
- Shuler KargiДокумент15 страницShuler Kargisureandhraindia67% (6)
- Biochem Enzyme KineticsДокумент53 страницыBiochem Enzyme KineticsJayvee Francisco67% (3)
- Separation Process Compilation of Problem SetДокумент55 страницSeparation Process Compilation of Problem Setjames100% (1)
- Enzyme KinecticsДокумент25 страницEnzyme KinecticsRhia80% (5)
- Gas AbДокумент14 страницGas AbJackielyn EugenioОценок пока нет
- Gas Solubility in Aqueous SolutionДокумент93 страницыGas Solubility in Aqueous SolutionArgie Adduru73% (11)
- Cell Kinetics and Fermenter Design Solution ManualДокумент5 страницCell Kinetics and Fermenter Design Solution ManualMarthy DayagОценок пока нет
- Chapter 3: Atomic and Ionic Arrangements: 3-1 SolutionДокумент52 страницыChapter 3: Atomic and Ionic Arrangements: 3-1 SolutionMarcos Jose100% (1)
- BIochem AssДокумент10 страницBIochem AssJomhel Callueng50% (2)
- COMPIДокумент129 страницCOMPIJeanne Roselle Dulatre CortezОценок пока нет
- Problems in Biochemical EngineeringДокумент22 страницыProblems in Biochemical EngineeringArrianne Jaye Mata50% (4)
- Basic Well Log Analysis - Introduction - Oct2013Документ35 страницBasic Well Log Analysis - Introduction - Oct2013Jorge PirelaОценок пока нет
- Optimize ethanol production from yeast in a continuous fermenterДокумент17 страницOptimize ethanol production from yeast in a continuous fermenterrhia81% (16)
- Biochemical Engineering Sample ProblemsДокумент5 страницBiochemical Engineering Sample ProblemsVan Vesper Dulliyao100% (1)
- CHE 511A Compilation 2 3 M F PDFДокумент164 страницыCHE 511A Compilation 2 3 M F PDFMaame Efua NeizerОценок пока нет
- 1097 2301 Univar E Detergents Nordic Brochure 4Документ12 страниц1097 2301 Univar E Detergents Nordic Brochure 4troity100% (2)
- Sample Problem #11Документ6 страницSample Problem #11Dozdi100% (4)
- LEC LE 3 Part 2 Stoichiometry of Growth and Product FormationДокумент27 страницLEC LE 3 Part 2 Stoichiometry of Growth and Product FormationRaymond Fuentes100% (1)
- Michaelis-Menten kinetics of enzyme reactionДокумент4 страницыMichaelis-Menten kinetics of enzyme reactionVan Vesper DulliyaoОценок пока нет
- Lecture 26-Problems On Cell Growth KineticsДокумент15 страницLecture 26-Problems On Cell Growth KineticsHemanth Peddavenkatappa GariОценок пока нет
- Cell Kinetics Chapter 6Документ56 страницCell Kinetics Chapter 6Sanjay Kumar73% (11)
- JJДокумент4 страницыJJEureca ParraОценок пока нет
- Bioche ProblemsДокумент5 страницBioche ProblemsTimothy Jones100% (1)
- Computer ApplicationsДокумент8 страницComputer Applicationsapi-3728602100% (1)
- Cell Kinetics and Fermenter DesignДокумент116 страницCell Kinetics and Fermenter DesignCriselda Oliva CarinoОценок пока нет
- CN3121 Homework Questions and Solutions (Chap1)Документ3 страницыCN3121 Homework Questions and Solutions (Chap1)Weng Hong WeiОценок пока нет
- 2 Cell Kinetics and Fermenter Design Part 2 DiscussionДокумент4 страницы2 Cell Kinetics and Fermenter Design Part 2 DiscussionEzekielОценок пока нет
- 5 Enzyme KineticsДокумент39 страниц5 Enzyme KineticsEbook Download100% (1)
- CHEMICAL ENGINEERING INSTRUMENTATION AND PROCESS CONTROL PROBLEMSДокумент4 страницыCHEMICAL ENGINEERING INSTRUMENTATION AND PROCESS CONTROL PROBLEMSMarco SarmientoОценок пока нет
- 14-Plant Design-Ans Key-Master FileДокумент22 страницы14-Plant Design-Ans Key-Master FilePaul Philip LabitoriaОценок пока нет
- Physical Chemistry: Answer KeyДокумент15 страницPhysical Chemistry: Answer Keyvishal110085Оценок пока нет
- Iit Jee 2012 Pet4 Solns p2Документ22 страницыIit Jee 2012 Pet4 Solns p2Ishita AggarwalОценок пока нет
- E 199 SolДокумент10 страницE 199 SolDavid Alemán SánchezОценок пока нет
- Introduction to Bioprocess Engineering Revision Questions and AnswersДокумент3 страницыIntroduction to Bioprocess Engineering Revision Questions and AnswersGia BooОценок пока нет
- Orca Share Media1521362143835Документ122 страницыOrca Share Media1521362143835Ana Lorraine DalilisОценок пока нет
- Solutions Manual Urban Drainage 2ndedДокумент17 страницSolutions Manual Urban Drainage 2ndedJamesBonn OlingaОценок пока нет
- Derive The Integrated Rate Equation Half-LifeДокумент7 страницDerive The Integrated Rate Equation Half-Lifeumut2000Оценок пока нет
- Department of Chemistry Faculty of Mathematics and Science State University of Padang 2014Документ9 страницDepartment of Chemistry Faculty of Mathematics and Science State University of Padang 2014Anelin OsiriknaОценок пока нет
- Chem II AP PacketДокумент4 страницыChem II AP PacketAmanda Rose DalyОценок пока нет
- Mach CC App Form Jun16 Eng 20160629Документ6 страницMach CC App Form Jun16 Eng 20160629ZulIzzamreeZolkepliОценок пока нет
- Hyundai AccentДокумент10 страницHyundai AccentVPrasarnth RaajОценок пока нет
- Job satisfaction in human resource managementДокумент102 страницыJob satisfaction in human resource managementMed ElbanadiОценок пока нет
- Windscreen Report Claim FormДокумент3 страницыWindscreen Report Claim FormVPrasarnth RaajОценок пока нет
- C19115I2901Документ1 страницаC19115I2901VPrasarnth RaajОценок пока нет
- 4 TH SOGCEDirectoryДокумент40 страниц4 TH SOGCEDirectoryVPrasarnth RaajОценок пока нет
- HP PDS Con EngДокумент3 страницыHP PDS Con EngVPrasarnth RaajОценок пока нет
- CarДокумент1 страницаCarVPrasarnth RaajОценок пока нет
- Finaldissertation NoRestrictionДокумент120 страницFinaldissertation NoRestrictionVPrasarnth RaajОценок пока нет
- Proton On The Road Price After Gst-20160215Документ1 страницаProton On The Road Price After Gst-20160215VPrasarnth RaajОценок пока нет
- ResignationДокумент1 страницаResignationAlan KhorОценок пока нет
- Map To CBJ OfficeДокумент1 страницаMap To CBJ OfficeVPrasarnth RaajОценок пока нет
- 2011 AccentДокумент8 страниц2011 AccentVPrasarnth RaajОценок пока нет
- LCAДокумент1 страницаLCAVPrasarnth RaajОценок пока нет
- Prototype Development To Test EOR Methods: Joana - Sanches@ist - Utl.ptДокумент10 страницPrototype Development To Test EOR Methods: Joana - Sanches@ist - Utl.ptVPrasarnth RaajОценок пока нет
- Jadual Waktu MOG Semester 2, Sesi 2014.2015Документ1 страницаJadual Waktu MOG Semester 2, Sesi 2014.2015VPrasarnth RaajОценок пока нет
- 308 - November 2015 Version 29Документ19 страниц308 - November 2015 Version 29VPrasarnth RaajОценок пока нет
- Pchardware StartecДокумент1 страницаPchardware StartecVPrasarnth RaajОценок пока нет
- All It DesktopДокумент4 страницыAll It DesktopNatasya WillmoreОценок пока нет
- Petroleum Home Assignment OutlineДокумент3 страницыPetroleum Home Assignment OutlineVPrasarnth RaajОценок пока нет
- Bioprocess QuizДокумент4 страницыBioprocess QuizVPrasarnth RaajОценок пока нет
- Rajni and ManagementДокумент12 страницRajni and ManagementVPrasarnth RaajОценок пока нет
- Milestone 5 Outline KC40203 2012Документ1 страницаMilestone 5 Outline KC40203 2012VPrasarnth RaajОценок пока нет
- List of Tables, Figures & Executive SummaryДокумент4 страницыList of Tables, Figures & Executive SummaryVPrasarnth RaajОценок пока нет
- Comparison of Different Collection Efficiency Models For Venturi ScrubbersДокумент10 страницComparison of Different Collection Efficiency Models For Venturi ScrubbersPassmore DubeОценок пока нет
- Packed Tower AbsorberДокумент58 страницPacked Tower AbsorberCelestino Montiel MaldonadoОценок пока нет
- Astm b680-1980Документ2 страницыAstm b680-1980Yavuz Solmaz100% (1)
- Bacteriological Properties and Health Related Biochemical Components of Fermented Fish Sauce - An Overview.Документ29 страницBacteriological Properties and Health Related Biochemical Components of Fermented Fish Sauce - An Overview.dizzydahlia97Оценок пока нет
- Chapter 03 AnnotДокумент4 страницыChapter 03 AnnotNur Amira Mardiana ZulkifliОценок пока нет
- Komatsu Powertrain Oil TO10Документ7 страницKomatsu Powertrain Oil TO10yoguijojoОценок пока нет
- SHS Physical Science Q1 SLM - 3Документ25 страницSHS Physical Science Q1 SLM - 3Adalee ColleenОценок пока нет
- COA CrosscarmДокумент1 страницаCOA CrosscarmSouheila MniОценок пока нет
- MSDS P Epoxy CureДокумент11 страницMSDS P Epoxy CureRUDDY ALFAN SUBAKTIОценок пока нет
- 06 Worksheet 2Документ4 страницы06 Worksheet 2Kinaston SkyОценок пока нет
- 6 9500 F Axera 7 240 PDFДокумент4 страницы6 9500 F Axera 7 240 PDFsegomezpОценок пока нет
- NG-018-D1-PNL-406701 - Rev06 UnsignedДокумент111 страницNG-018-D1-PNL-406701 - Rev06 UnsignedFolarin AyodejiОценок пока нет
- Improvements of Foundation RocksДокумент2 страницыImprovements of Foundation RocksSteven Joseph Incio100% (1)
- 72 Had DadДокумент6 страниц72 Had DadthestupiddogОценок пока нет
- What Is WeldingДокумент4 страницыWhat Is WeldingAnonymous hBBam1nОценок пока нет
- Polished Rod Clamp Brochure Rev5Документ2 страницыPolished Rod Clamp Brochure Rev5Pedro Antonio Mejia SuarezОценок пока нет
- Soils of Karnataka-361Документ10 страницSoils of Karnataka-361naturewisdom100% (3)
- Transpiration: Prepared By: Concepcion, Ada - Trinidad, Lester - Tolon, ChristianДокумент17 страницTranspiration: Prepared By: Concepcion, Ada - Trinidad, Lester - Tolon, ChristiannimhaОценок пока нет
- Ammonium 2520sulphate Material 2520balance.Документ9 страницAmmonium 2520sulphate Material 2520balance.AgadmatorОценок пока нет
- Tài liệu ôn tập tiếng anh 4Документ7 страницTài liệu ôn tập tiếng anh 4Ngọc AmiiОценок пока нет
- Avoid Costly Materials mistakes-CEДокумент23 страницыAvoid Costly Materials mistakes-CEManish542Оценок пока нет
- Davey XS250HG ManualДокумент2 страницыDavey XS250HG Manualcoolestkiwi100% (1)
- Hotel FireДокумент94 страницыHotel FireCoara OvidiuОценок пока нет
- Phosphorus, Reactive, TNT, PhosVer 3 Method 8048, 02-2009, 9th EdДокумент7 страницPhosphorus, Reactive, TNT, PhosVer 3 Method 8048, 02-2009, 9th EdRaúl BatistaОценок пока нет