Академический Документы
Профессиональный Документы
Культура Документы
Endogenous Antimicrobial Peptides and Skin Infections in Atopic Dermatitis
Загружено:
Taqwa NindaОригинальное название
Авторское право
Доступные форматы
Поделиться этим документом
Поделиться или встроить документ
Этот документ был вам полезен?
Это неприемлемый материал?
Пожаловаться на этот документАвторское право:
Доступные форматы
Endogenous Antimicrobial Peptides and Skin Infections in Atopic Dermatitis
Загружено:
Taqwa NindaАвторское право:
Доступные форматы
ANT IMICROB IA L PEP TID ES IN ATOPIC D ERMATITIS
ENDOGENOUS ANTIMICROBIAL PEPTIDES AND SKIN INFECTIONS
IN ATOPIC DERMATITIS
PECK Y. ONG, M.D., TAKAAKI OHTAKE, M.D., PH.D., CORINNE BRANDT, B.S., IAN STRICKLAND, PH.D.,
MARK BOGUNIEWICZ, M.D., TOMAS GANZ, M.D., PH.D., RICHARD L. GALLO, M.D., PH.D.,
AND DONALD Y.M. LEUNG, M.D., PH.D.
ABSTRACT
Background The innate immune system of human
skin contains antimicrobial peptides known as cathelicidins (LL-37) and b -defensins. In normal skin these
peptides are negligible, but they accumulate in skin
affected by inflammatory diseases such as psoriasis.
We compared the levels of expression of LL-37 and
human b-defensin 2 (HBD-2) in inflamed skin from
patients with atopic dermatitis and from those with
psoriasis.
Methods The expression of LL-37 and HBD-2 protein in skin-biopsy specimens from patients with psoriasis, patients with atopic dermatitis, and normal
subjects was determined by immunohistochemical
analysis. The amount of antimicrobial peptides in extracts of skin samples was also analyzed by immunodot blot analysis (for LL-37) and Western blot analysis
(for HBD-2). Quantitative, real-time reverse-transcriptasepolymerase-chain-reaction (RT-PCR) assays were
used to confirm the relative expression of HBD-2 and
LL-37 messenger RNA (mRNA) in the skin-biopsy
specimens. These peptides were also tested for antimicrobial activity against Staphylococcus aureus with
the use of a colony-forming assay.
Results Immunohistochemical analysis confirmed
the presence of abundant LL-37 and HBD-2 in the superficial epidermis of all patients with psoriasis. In
comparison, immunostaining for these peptides was
significantly decreased in acute and chronic lesions
from patients with atopic dermatitis (P=0.006 and
P=0.03, respectively). These results were confirmed
by immunodot blot and Western blot analyses. Realtime RT-PCR showed significantly lower expression
of HBD-2 mRNA and LL-37 mRNA in atopic lesions
than in psoriatic lesions (P=0.009 and P=0.02, respectively). The combination of LL-37 and HBD-2 showed
synergistic antimicrobial activity by effectively killing
S. aureus.
Conclusions A deficiency in the expression of antimicrobial peptides may account for the susceptibility of patients with atopic dermatitis to skin infection
with S. aureus. (N Engl J Med 2002;347:1151-60.)
HE skins first line of defense against invasion by microbial agents is the stratum corneum, a nonviable, desiccated layer of the
epidermis.1 However, this physical barrier
is susceptible to injuries that allow the entry of opportunistic microbial agents into the skin. The innate
immune system can immediately respond to this intrusion by helping to prevent further invasion. This
immune response includes phagocytosis by neutrophils and macrophages and their production of reactive oxygen intermediates that kill microbial agents.2
A number of endogenous antimicrobial peptides
have been shown to play an integral part in innate immunity.3 Two major classes of peptides in mammalian
skin, b-defensins4,5 and cathelicidins,6,7 have antimicrobial activity against bacterial, fungal, and viral pathogens.4,7,9 These peptides, which are produced by keratinocytes in the skin,4,7,8 disrupt the membrane of
the target microbe or penetrate the microbial membrane, interfering with intracellular functions.10 The
expression of some of these peptides, such as human
b-defensin 1 (HBD-1), is constitutive,8 whereas the
expression of others, including human b-defensin 2
(HBD-2) and LL-37, a cathelicidin, is triggered by
injury or inflammation of the skin.4,7 Animal models
have shown that the expression or activation of antimicrobial peptides is essential for the ability of skin to
resist bacterial infection.11,12
Atopic dermatitis, a chronic inflammatory skin disease frequently found in families with asthma and allergic rhinitis,13 is complicated by recurrent infections
of skin lesions by bacterial, viral, and fungal pathogens.14 About 30 percent of patients with atopic dermatitis have bacterial or viral infections of the skin, as
compared with only 7 percent of patients with psoriasis,15 even though both diseases are characterized
by a defective skin barrier.16 We compared the expression of HBD-2 and LL-37 in skin lesions from pa-
Copyright 2002 Massachusetts Medical Society.
From the Division of Allergy and Immunology, Department of Pediatrics, National Jewish Medical and Research Center, Denver (P.Y.O., I.S.,
M.B., D.Y.M.L.); the Department of Pediatrics, University of Colorado
Health Sciences Center, Denver (M.B., D.Y.M.L.); the Division of Dermatology, Department of Medicine and Pediatrics, University of California,
San Diego, and the Veterans Affairs San Diego Health Care System, San
Diego (T.O., C.B., R.L.G.); and the Division of Pulmonary and Critical
Care Medicine, Departments of Medicine and Pathology, School of Medicine, University of California, Los Angeles (T.G.). Address reprint requests
to Dr. Leung at the National Jewish Medical and Research Center, Department of Pediatrics, Rm. K926, 1400 Jackson St., Denver, CO 80206.
N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org 1151
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
The Ne w E n g l a nd Jo u r n a l o f Me d ic i ne
tients with atopic dermatitis with their expression in
psoriatic lesions and normal skin, using immunohistochemical staining, Western and immunodot blotting,
and a quantitative, real-time reverse-transcriptase
polymerase-chain-reaction (RT-PCR) assay. We also
examined the capacity of tumor necrosis factor a
(TNF-a) to induce HBD-2 expression in a human keratinocyte cell line after treatment with interleukin-4
and interleukin-13, cytokines that are abundant in the
skin of patients with atopic dermatitis.13 Finally, we
evaluated the combined antimicrobial effects of LL-37
and HBD-2 on Staphylococcus aureus.
METHODS
Patients
The study participants included 8 patients with moderate-tosevere atopic dermatitis (mean age, 33 years; extent of skin involvement, 20 to 60 percent), 11 patients with psoriasis (mean age, 38
years; extent of skin involvement, 15 to 40 percent), and 6 healthy
persons (mean age, 38 years). None of the patients had received
systemic corticosteroids previously, and none had received topical
corticosteroids for at least one week before enrollment. The study
was approved by the institutional review board at National Jewish
Medical and Research Center, in Denver; all patients and normal
subjects gave written informed consent.
Punch biopsies were performed, with 2-mm samples obtained
from erythematous lesions that were less than three days old (acute
atopic dermatitis), lichenified lesions that were more than two weeks
old (chronic atopic dermatitis), psoriatic lesions, and normal skin.
The skin samples were immediately frozen at 70C for immunohistochemical studies or Western and immunodot blot analyses.
Immunohistochemical Staining
For immunostaining of HBD-2, frozen 5-m skin-tissue sections
were fixed in acetone for 10 minutes and then incubated with 10
percent nonimmune goat serum (Zymed) for 10 minutes. Blocking serum was removed, and the sections were stained with anti
HBD-2 rabbit polyclonal antibody (Peptide Institute) at a 1:50 dilution (vol/vol) in phosphate-buffered saline for one hour at room
temperature in a humid atmosphere. The slides were rinsed twice
with phosphate-buffered saline and incubated with a biotinylated
goat antirabbit secondary antibody (Zymed) at room temperature
for 30 minutes, rinsed twice with phosphate-buffered saline, and incubated with a mixture of avidin and biotinylated horseradish peroxidase (1:1 vol/vol, Dako) for an additional 30 minutes. The reaction was developed with the use of amino-ethyl-carbazole single
solution (Zymed) for 10 minutes and then counterstained with hematoxylin. Negative controls were established with the use of nonimmune isotype antibodies and preincubation of the antiHBD-2
antibodies with a synthetic HBD-2 peptide (Peptide Institute) to
ensure the binding specificity of the antiHBD-2 antibodies. All
slides were coded before the samples were evaluated so that the identity of the study subjects was not revealed. The intensity of the
immunostaining was graded with the use of microscopy on a scale
from 0 to 3, with 0 indicating no staining and 3 the most intense
staining.
For immunostaining of LL-37, frozen skin sections were treated
similarly at first and then incubated with rabbit antiLL-37 antibody, as previously described,17 in phosphate-buffered saline and
0.1 percent bovine serum albumin. The sections were washed in
phosphate-buffered saline and stained with goat antirabbit horseradish peroxidase (Vectastain Elite ABC Rabbit kit, Vector Laboratories) and diaminobenzidine substrate (Sigma) according to the
manufacturers instructions. The sections were counterstained with
hematoxylin. The specificity of the primary antibody reaction was
confirmed in separate experiments by adsorption of either anti
LL-37 antibody with excess amounts of the synthetic peptide. The
specificity of the secondary antibody reaction and of the immunostaining reagents was confirmed by routine use of rabbit nonimmune serum.
Measurement of HBD-2 and LL-37 Peptides
Skin-biopsy samples were weighed and then homogenized for
10 minutes with a loose-fitting Dounce homogenizer in 1 ml of
1 M hydrogen chloride and 1 percent trifluoroacetic acid on ice. Homogenized tissues in solution were then rotated overnight at 4C.
After centrifugation for 20 minutes at 14,000 rpm at 4C, supernatants were transferred to new tubes and lyophilized completely. The
resulting protein pellets were dissolved with distilled water to obtain a final protein concentration of 0.1 mg of tissue per microliter.
For the measurement of HBD-2, 15 l of each sample was lyophilized again and resuspended in sodium dodecyl sulfate buffer
overnight at 4C. Samples and known amounts of HBD-2 for use
as standards were then boiled for five minutes and loaded onto a
16.5 percent sodium dodecyl sulfatetricine polyacrylamide gel.
The samples were placed on ImmobilonPSQ membranes (Millipore) for one hour at 0.18 mA in 0.05 M sodium borate, pH 9.0,
with 20 percent methanol and 0.05 percent sodium dodecyl sulfate. Blots were fixed for 30 minutes with 0.5 percent glutaraldehyde
in TRIS-buffered saline (500 mM sodium chloride and 20 mM
TRIS, pH 7.5), blocked for 30 minutes in 0.75 percent Blotto
(nonfat powdered milk) in phosphate-buffered saline (0.9 percent
sodium chloride and 10 mM sodium phosphate buffer, pH 7.4),
then incubated for 18 hours in a 1:500 dilution of rabbit anti
HBD-2 serum in antibody dilution buffer (0.25 percent Blotto
in phosphate-buffered saline containing 0.01 percent thimerosal as
a preservative). The blots were washed in 0.1 percent bovine serum
albumin in Blotto TRIS-buffered saline (0.9 percent sodium chloride and 20 mM TRIShydrogen chloride, pH 4.5 to 5.0) three
times for 10 minutes each time. The membranes were incubated
in a 1:2000 dilution of alkaline phosphataseconjugated goat antirabbit IgG in antibody dilution buffer for one hour, then washed
three times as before and developed in alkaline phosphatase development solution (bromochloroindolyl phosphatenitroblue tetrazolium).
For the measurement of LL-37, immunodot blot analysis was
performed. Positive controls and standard curves were generated
with a synthetic LL-37 peptide (C-terminal, 18-mer fragment; amino acid sequence, CZQPIKDFLRNLVPRTES; molecular weight,
2204). The peptide was diluted serially from 50 to 1600 nM;
100 l of each peptide standard and 5 l of each sample (equivalent
to 0.5 mg of tissue), extracted in a manner identical to that for
HBD-2 measurements, were dotted in triplicate on nitrocellulose
membrane. The membrane was blocked with 10 percent nonfat
dry milk (Bio-Rad) and 0.1 percent Tween 20 in TRIS-buffered saline at room temperature for three hours and then incubated at
4C overnight with rabbit antiLL-37 antibody diluted to 1:2500
in the blocking solution. The membrane was then incubated at
room temperature for two hours with goat antirabbit antibody
horseradish peroxidase (Dako) that was diluted to 1:4000 in
blocking solution. Western Lightning chemiluminescence reagent
(PerkinElmer Life Sciences) was used to detect electro-chemiluminescence. A low-light imaging system (ChemiImager 4400, Alpha Innotech) was used for densitometrical analysis. Signal detected
by dot blot analysis was confirmed in some experiments by Western blot analysis, which identified full-length LL-37 at 18 kD. For
quantification of both LL-37 and HBD-2, the signal intensity of
tissue extracts was directly compared with that of a simultaneously
prepared standard consisting of known amounts of each synthetic
peptide. The concentration of antimicrobial peptide in the original
skin-biopsy sample was then estimated by dividing the experimentally determined mass of antimicrobial peptide from the skin-biopsy
1152 N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
ANT IMICROBIA L PEP TID ES IN ATOPIC D ERMATITIS
sample by the epidermal volume. The epidermal volume was estimated by assuming an epidermal thickness of 0.1 mm; thus, the
volume of epidermis in a 2-mm punch-biopsy specimen was
3.14101 mm3.
RT-PCR Analysis of HBD-2 and LL-37 Messenger RNA
Total RNA was isolated from 2-mm skin-biopsy samples from
the subjects with the use of TRI Reagent (Sigma) according to the
manufacturers protocol. The primers and probe for HBD-2 were
designed with the use of Prism 7700 sequence-detection software
(Primer Express, PerkinElmer Applied Biosystems). The primer
sequences were 5'TCCTCTTCTCGTTCCTCTTCATATTC3' (forward primer) and 5'TTAAGGCAGGTAACAGGATCGC3' (reverse
primer). The sequence for the probe (TaqMan, PerkinElmer) was
5'ACCACCAAAAACACCTGGAAGAGGCA3'; the 5' end was labeled with 6-carboxyfluorescein, and the 3' end was labeled with
6-carboxytetramethylrhodamine.
Amplification reactions were performed with the use of a sequence detector (ABI Prism 7700, PerkinElmer Applied Biosystems) in MicroAmp optical tubes (PerkinElmer Applied Biosystems) in a 25-l mixture containing 8 percent glycerol, 1 TaqMan
buffer A (500 mM potassium chloride, 100 mM TRIS-hydrogen
chloride, 0.1 M EDTA, and 600 nM passive reference dye ROX,
pH 8.3, at room temperature), 300 M each of deoxyadenosine
triphosphate, deoxyguanosine triphosphate, and deoxycytidine triphosphate, and 600 M deoxyuridine triphosphate; 5.5 mM magnesium chloride; 900 nM forward primer; 900 nM reverse primer;
200 nM probe; 0.625 U of AmpliTaq Gold DNA Polymerase (PerkinElmer); 6.25 U of Moloney murine leukemia virus reverse
transcriptase (Life Technologies); 10 U of RNasin ribonuclease inhibitor (Promega); and the template RNA. The RT assay was performed at 48C for 30 minutes, followed by activation of TaqGold
at 95C for 10 minutes. Forty cycles of amplification were then
performed at 95C for 15 seconds and at 60C for 1 minute.
After amplification, real-time data acquisition and analysis were
performed. The fluorescence data were expressed as normalized reporter signal, calculated by dividing the amount of reporter signal
by the amount of passive reference signal, or as the change in the
normal reporter signal, calculated as the amount of normalized reporter signal minus the amount of reporter signal before PCR. The
detection threshold was set above the mean base-line value for fluorescence determined on the basis of the first 15 cycles. Amplification reactions in which the intensity of fluorescence exceeded the
threshold were defined as positive reactions. The threshold cycle
(Ct) is the PCR cycle at which an increase in reporter signal above
the base-line signal can first be detected. A standard curve was
generated with the use of the fluorescence data from the 10-fold
serial dilutions of total RNA from a skin-biopsy sample from a patient with psoriasis. This curve was used to calculate the relative
amounts of HBD-2 in test samples. Quantities of HBD-2 in test
samples were normalized to the corresponding 18S ribosomal RNA
(rRNA) (P/N 4308310, PerkinElmer Applied Biosystems).
The real-time, quantitative PCR assay for LL-37 was performed
with the use of a sequence-detection system (GeneAmp 5700, PerkinElmer). Classic 18S primers (Ambion) were used to amplify 18S
rRNA. The primer sequences used to amplify LL-37 complementary DNA (cDNA) were 5'GCAGTCACCAGAGGATTGTGAC3'
(forward primer) and 5'CACCGCTTCACCAGCCC3' (reverse
primer). For the PCR assay of LL-37, 1.2 l of RT reaction was
used; 1.2 l of 200-fold diluted RT reaction was used for PCR of
18S rRNA. Amplification reactions were performed in a final total
volume of 25.7 l containing SYBR Green PCR Master Mix (Applied Biosystems), 10 M primer, and nuclease-free water. The assay was performed at 50C for 2 minutes and at 95C for 10 minutes. Forty cycles of amplification were then performed at 94C for
15 seconds and at 60C for 1 minute. The results were analyzed
with the use of the comparative Ct method. This method is based
on the assumption that the target (LL-37) and reference (18S)
primers amplify with the same efficiency within a given range of
initial messenger RNA (mRNA) concentrations. To test this assumption, cDNA was made from total RNA of a psoriasis sample
at the initial total RNA concentrations of 1000, 500, 100, 50, and
10 ng. Real-time PCR was performed in triplicate for each starting
concentration, and the change in Ct (Ct) was calculated. For
each starting concentration of total RNA, Ct=CtR CtG, where
CtR denotes the reference Ct and CtG the target Ct. This value was
plotted against the log of the initial total RNA concentration. The
efficiencies of gene amplification were close enough that a correction factor over the specified range was not required if the slope was
less than or equal to 0.1. We calculated the Ct for samples as
follows: Ct= CtE CtB, where CtE denotes the experimental
Ct and CtB the base-line Ct. The base-line sample was a normal
control sample, which had low expression of LL-37 mRNA. The relative expression was calculated as 2Ct in order to account for the
exponential amplification of the PCR reaction.
Cell Culture
HaCat cells, a human keratinocyte cell line, were grown in Dulbeccos Modified Eagles medium (Cellgro), supplemented with 10
percent fetal-calf serum (Gemini) and 1 percent of each of the following: 200 mM L-glutamine, minimal essential medium (MEM)
with nonessential amino acids (GIBCO), MEM Vitamins Solution
(Life Technologies), and penicillinstreptomycin. To study the effects of interleukin-4 and interleukin-13 on HBD-2 mRNA expression in HaCat cells, 5105 cells per milliliter were incubated in
control medium, 20 ng of TNF-a per milliliter alone or 20 ng of
TNF-a per milliliter plus 50 ng of interleukin-4 per milliliter, alone
or in combination with 50 ng of interleukin-13 per milliliter, for
24 hours. The cells were then washed once and homogenized in
TRI reagent by repeated pipetting. Total RNA was isolated according to the manufacturers protocol.
Measurement of Antimicrobial Activity
LL-37 and HBD-2 peptides were synthesized as previously described.17,18 The antimicrobial activity of the peptides was determined with the use of a solution colony-forming unit (CFU) assay.
Wild-type S. aureus isolated from patients with atopic dermatitis was
assessed. Bacteria were grown to a concentration at which they were
multiplying exponentially (mid log phase; A600 1.5=8.010 9
CFU per milliliter) in Todd Hewitt broth medium (Sigma). Bacteria were washed twice with 10 mM sodium phosphate buffer (pH
7.4) containing 0.1 percent (wt/vol) Todd Hewitt broth and were
diluted to a final concentration of 2.0107 cells per milliliter in the
same buffer. Ten microliters of this bacterial suspension was then
placed in round-bottom, 96-well cell-culture plates and incubated
at 37C and 100 percent humidity for two hours with various concentrations of LL-37, HBD-2, or both. The cytotoxic activity and
minimal bactericidal activity of LL-37 and HBD-2 were analyzed
by plating serial dilutions of the incubation mixture in triplicate
on Todd Hewitt broth agar plates and determining the CFU the
following day. The percentage of killed bacteria was expressed as
[1(CFU after peptide incubation)(CFU after control incubation)]100.
Statistical Analysis
The nonparametric median test from the JMP4 statistical package19 was used to determine significant differences. We constrained
experimentwise error rates to the standard alpha level of 0.05 by
the Bonferroni method; the alpha level was 0.025 for a comparison
of two means and 0.017 for a comparison of three means.
RESULTS
Figure 1 shows the immunostaining of HBD-2 and
LL-37 in skin-biopsy samples. The samples from pso-
N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org 1153
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
The Ne w E n g l a nd Jo u r n a l o f Me d ic i ne
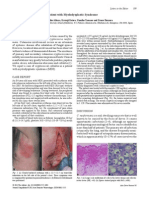
Figure 1. Immunostaining for HBD-2 (Panel A)
and LL-37 (Panel B) in Frozen Skin Sections
from Patients with Psoriasis or Atopic Dermatitis (AD) and Normal Subjects.
Immunostaining for HBD-2 and LL-37 was
more intense in the psoriatic lesions than in
the atopic lesions or normal skin. The arrows
in Panels A and B indicate immune reactivity
to HBD-2 and LL-37, respectively.
Psoriasis
Normal
Chronic AD
Acute AD
Psoriasis
Normal
Chronic AD
Acute AD
1154 N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
ANT IMICROB IA L PEP TID ES IN ATOPIC D ERMATITIS
riatic lesions had much more intense staining for both
HBD-2 and LL-37 than the samples from acute or
chronic atopic lesions or normal skin. Figure 2 shows
the composite data on HBD-2 immunostaining for
all the skin-biopsy samples. The intensity of the immunostaining of samples from acute atopic lesions did
not differ significantly from that of chronic atopic lesions or normal skin. Psoriatic lesions, however, had
significantly more intense immunostaining than acute
atopic lesions (P=0.006), chronic atopic lesions (P=
0.03), or normal skin (P=0.02). The results of immunostaining for LL-37 in the samples of normal skin
and psoriatic lesions were consistent with a previous
study showing that psoriatic lesions have greater expression of LL-37 than normal skin.7 The inflammatory skin lesions from patients with atopic dermatitis
had markedly lower expression of LL-37 than the skin
lesions from patients with psoriasis. The biggest difference was in the region of the granular cell layer
and stratum corneum, which was relatively devoid of
LL-37 in the atopic lesions but had the highest expression of LL-37 in the psoriatic lesions. The absence of
HBD-2 and LL-37 in the normal skin also confirms
previous reports that these antimicrobial peptides are
not normally produced but are up-regulated under inflammatory conditions.4,7
For further quantification of the relative amounts of
antimicrobial peptides in skin lesions, 2-mm biopsy
specimens were acid-extracted, and the total amount
of LL-37 and HBD-2 was measured (Fig. 3). Sufficient skin samples were available for analysis from only
five patients with psoriasis and six with atopic dermatitis. Western blotting, as compared with a standard curve of recombinant HBD-2, showed that the
amount of HBD-2 present in the psoriatic lesions
ranged from 2.3 to 157 M (median, 20), whereas no
HBD-2 was detectable in the atopic lesions (Fig. 3A).
The results of quantification of LL-37 in skin-biopsy
samples with the use of immunodot blot analysis were
similar, with an abundance of LL-37 in the psoriatic
lesions (median, 304 M; range, 0 to 1605); only one
of the six atopic lesions had detectable LL-37 (Fig.
3B). The single positive sample from a patient with
atopic dermatitis was from a chronic lesion, probably
reflecting the contribution of LL-37 from an inflammatory-cell infiltrate.
The difference in the expression of HBD-2 and
LL-37 protein between atopic lesions and psoriatic
P=0.006
P=0.03
P=0.02
Intensity of Staining
0
Acute
Chronic
Atopic Dermatitis
Psoriasis
Normal
Figure 2. Intensity of Immunostaining for HBD-2 in Skin from Patients with Atopic Dermatitis, Patients with Psoriasis, and Normal Subjects.
The intensity of the staining was graded visually on a scale from 0 (no staining) to 3 (the
most intense staining). There was no staining in any of the skin-biopsy samples from normal subjects.
N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org 1155
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
The Ne w E n g l a nd Jo u r n a l o f Me d ic i ne
P=0.002
HBD-2 (mM)
150
100
50
0
Acute
Chronic
Atopic Dermatitis
Psoriasis
P=0.04
LL-37 (mM)
3000
2000
1000
0
Acute
Chronic
Atopic Dermatitis
Psoriasis
Figure 3. Quantification of HBD-2 (Panel A) and LL-37 (Panel B)
in Epidermis of Atopic and Psoriatic Skin Lesions.
Data for HDB-2 are based on Western blot analysis, and data
for LL-37 on immunodot blot analysis.
lesions was further confirmed by the measurement of
mRNA with the use of real-time RT-PCR. As shown
in Figure 4, the expression of HBD-2 mRNA was significantly lower in acute atopic lesions (median, 98 pg
per nanogram of rRNA) and chronic atopic lesions
(median, 3268 pg per nanogram of rRNA) than in
psoriatic lesions (median, 31,660 pg per nanogram
of rRNA; P=0.009 for both comparisons). Real-time
RT-PCR analyses of LL-37 mRNA showed similar results, with significantly lower expression of LL-37 pep-
tide in acute and chronic atopic lesions than in psoriatic lesions (P=0.02 for both comparisons) (Fig. 4).
Previous studies have shown that LL-37 and
HBD-2 are greatly increased in patients with inflammatory skin conditions.5-7 Therefore, the diminished
expression of the peptides in lesions from patients with
atopic dermatitis was an unexpected observation. To
determine the mechanism underlying the reduced expression of antimicrobial peptides in atopic dermatitis,
we studied the effects of two cytokines (interleukin-4
and interleukin-13) that are elevated in atopic dermatitis but not in psoriasis.13 Figure 5 shows the effects of
interleukin-4 and interleukin-13 on TNF-ainduced
expression of HBD-2 in HaCat cells. Interleukin-4
alone or in combination with interleukin-13 significantly suppressed the up-regulation of HBD-2 mRNA
by TNF-a (P=0.04). In normal human skin, both
interleukin-4 and interleukin-13 suppressed TNF-a
induced HBD-2 mRNA expression in these explants
(data not shown). Since atopic lesions are characterized by the predominance of interleukin-4 and interleukin-13 expression, the suppression of HBD-2
mRNA by these cytokines may account for the low
expression of HBD-2 in these lesions.
The relative absence of HBD-2 and LL-37 in the
epidermis in atopic lesions, as compared with psoriatic lesions, suggested that the lack of these antimicrobial peptides in patients with atopic dermatitis might
contribute to their susceptibility to bacterial colonization and infection. To investigate this possibility, we
evaluated the susceptibility of clinical isolates of S. aureus to cytotoxic activity by both LL-37 and HBD-2.
Alone, LL-37 showed strong activity (Fig. 6A), whereas HBD-2 was much less potent (minimal bactericidal
concentration, >160 M). No significant difference
in susceptibility to these antimicrobial peptides was
detected in clinical isolates, as compared with wildtype S. aureus (American Type Culture Collection
number 25923), but all the isolates were more resistant than defensin-sensitive strains lacking cell-wall
charge modifications that confer resistance.20 The relative resistance of S. aureus isolates from patients with
atopic dermatitis argues against the reversion of the
bacteria to less resistant forms in this disorder. When
these resistant strains of S. aureus were exposed to
antimicrobial peptides expressed in psoriatic skin, however, the presence of both peptides enhanced their cytotoxic activity (Fig. 6). A combination of HBD-2 (at
80 M or 160 M) and LL-37 (at 4 M) killed significantly more S. aureus organisms than did HBD-2
alone.
DISCUSSION
Atopic dermatitis is a very common skin disease
known to be associated with a high prevalence of skin
infections, particularly with S. aureus.13 The density
1156 N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
ANT IMICROB IA L PEP TID ES IN ATOPIC D ERMATITIS
P=0.009
P=0.02
P=0.009
P=0.02
50
80,000
Ratio of LL-37 mRNA to rRNA
HBD-2 mRNA (pg/ng of rRNA)
10
50,000
40,000
30,000
20,000
10,000
9
8
7
6
5
4
3
2
1
0
Acute
Chronic
Atopic Dermatitis
Acute
Psoriasis
Chronic
Atopic Dermatitis
Psoriasis
Figure 4. HBD-2 and LL-37 Messenger RNA (mRNA) in Atopic and Psoriatic Skin Lesions.
Data are based on real-time reverse-transcriptasepolymerase-chain-reaction analyses of HBD-2 (Panel A) and LL-37 (Panel B). The
abbreviation rRNA denotes ribosomal RNA.
P=0.04
P=0.04
HBD-2 mRNA (pg/ng of rRNA)
15
10
Medium
TNF-a
Interleukin-4
TNF-a
and
Interleukin-4
Interleukin-13
TNF-a,
Interleukin-4,
and
Interleukin-13
Figure 5. Effects of Interleukin-4 and Interleukin-13 on Tumor Necrosis Factor a (TNF-a)Induced HBD-2 Expression in HaCat Cells.
The concentration of TNF-a was 20 ng per milliliter, and the concentrations of interleukin-4 and interleukin-13 were 50 ng per milliliter
each. The data are based on a total of three experiments. The bars indicate standard deviations, and rRNA denotes ribosomal RNA.
N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org 1157
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
The Ne w E n g l a nd Jo u r n a l o f Me d ic i ne
Cytotoxic Activity (%)
150
100
50
0
50
100
16
32
LL-37 (mM)
C
B
P=0.03
Cytotoxic Activity (%)
40
20
20
20
20
40
40
60
60
80
P=0.02
40
80
LL-37 (mM)
Figure 6. Cytotoxic Activity of HBD-2 and LL-37 against Wild-Type Staphylococcus aureus.
Wild-type S. aureus (2.0107 per milliliter) was incubated with increasing concentrations of LL-37
alone (Panel A) or LL-37 combined with HBD-2 at a concentration of 80 m (Panel B) or 160 m (Panel C).
The I bars denote standard deviations.
of S. aureus in inflamed atopic lesions without clinical
superinfection can be as high as 107 colony-forming
units per square centimeter of lesional skin. The important role of S. aureus is supported by the observation that even in patients with atopic dermatitis who
do not have superinfection, the severity of the skin
disease is reduced by treatment with a combination
of antistaphylococcal antibiotics and topical corticosteroids.21
Since S. aureus infection is a frequent trigger for the
exacerbation of skin disease, there has been considerable interest in the mechanisms underlying the increased colonization of atopic skin lesions with S. aureus.22 Atopic skin has significantly greater binding
affinity for S. aureus than nonatopic or psoriatic skin,
probably as the result of underlying inflammation.23
This view is supported by the observation that treat-
ment with topical glucocorticoids or tacrolimus reduces S. aureus counts on atopic skin.24,25 Indeed, a recent
study has shown that the number of S. aureus organisms that bind to inflammatory skin lesions is significantly higher when the inflammation is mediated by
type 2 helper T cells than when it is mediated by type
1 helper T cells.26
The increased avidity of S. aureus for atopic skin,
however, can account only for a several-fold increase
in S. aureus on atopic skin.23 Examination of skinbiopsy samples from patients with atopic dermatitis
has shown that S. aureus grows in colonies in the upper layers of the epidermis between keratinocytes.27
This suggests that an exponential increase in S. aureus
could result from failure of the innate immune defense
system of atopic skin to restrict the growth of the organisms. Naturally occurring antimicrobial peptides
1158 N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
ANT IMICROB IA L PEP TID ES IN ATOPIC D ERMATITIS
are a critical component of this innate immune system
that have been shown to provide mammalian skin with
resistance to bacterial infection.11 The antimicrobial
peptides HBD-2 and LL-37 are normally produced
by keratinocytes in response to inflammatory stimuli
such as psoriasis or injury.7 Consistent with the concept that the activity of keratinocytes in atopic dermatitis differs from that in psoriasis, recent observations
indicate that in atopic dermatitis, but not psoriasis, keratinocytes produce a distinct profile of chemokines
that promote the influx of eosinophils and type 2 helper T cells into the skin, whereas psoriatic keratinocytes
promote infiltration by neutrophils and type 1 helper
T cells.28
In our study, several independent approaches
demonstrated that at both the protein level and the
mRNA level, LL-37 and HBD-2 are deficient in skin
lesions from patients with atopic dermatitis. We also
demonstrated that the combination of HBD-2 and
LL-37 at the concentrations found in psoriatic lesions,
but not atopic lesions, was sufficient to kill S. aureus.
These antimicrobial peptides have activity not only
against bacteria but also against fungi and viruses.
Thus, our observations may explain the increased susceptibility of patients with atopic dermatitis to fungal
and viral infections such as herpes simplex and molluscum contagiosum.
We also investigated the potential mechanism (or
mechanisms) for the reduced expression of antimicrobial peptides such as HBD-2. Since it is well established that atopic skin lesions are associated with increased expression of interleukin-4 and interleukin-13,
we explored the possibility that these type 2 helper
cytokines may inhibit HBD-2 gene expression. Indeed, we found that the combination of interleukin-4
and interleukin-13 was highly effective in inhibiting
HBD-2 gene expression. Thus, type 2 helper cytokines
account not only for the high IgE concentrations and
eosinophilia in patients with atopic dermatitis but also
for their increased susceptibility to skin infections.
In summary, our study showed that inflammatory
skin lesions from patients with atopic dermatitis had
significantly lower concentrations of HBD-2 and
LL-37 than skin lesions from patients with psoriasis.
The inhibitory effects of interleukin-4 and interleukin-13, both of which are abundant in atopic skin, may
account for this finding. Furthermore, the low concentrations of HBD-2 and LL-37 in the patients with
atopic dermatitis were unable to kill S. aureus. Therefore, a deficiency in the expression of inflammationinduced antimicrobial peptides may explain the susceptibility of patients with atopic dermatitis to skin
infections. Our findings demonstrate the presence of
skin-localized immunodeficiency in atopic dermatitis
and the correlation between deficient expression of
antimicrobial peptides and infection. These findings
point to the role of antimicrobial-peptide function in
clinical disease and highlight the importance of considering the interaction between the innate and adaptive immune systems.
Supported in part by grants from the National Institutes of Health
(HL36577, AR41256, and HL37260), the Division of Research Resources
(Clinical Research Center grant MO1 RR00051), and the University of
Colorado Cancer Center to Dr. Leung; a National Research Service Award
(T32 AI 07365) and a Fujisawa Health Care Allergic Skin Diseases Award
to Dr. Ong; a Veterans Affairs Merit Review Award and grants from the
Stern Foundation and the National Institutes of Health (AI48176) to Dr.
Gallo; and a grant from the National Institutes of Health (HL46809) to
Dr. Ganz.
We are indebted to Birgitta Agerberth for the gift of antiLL-37
antibody, to Dr. Bryan R. Hauge and Umarani Pugazhenthi at the
University of Colorado Cancer Center Core Facility for assistance
with the real time RT-PCR analyses of HBD-2, to Maureen Sandoval for assistance with the preparation of the manuscript, to Dr. James
Murphy for assistance with the statistical analyses, and to Dr. Hal
Jenkins for his critical reading of the manuscript.
REFERENCES
1. Jackson SM, Elias PM. Skin as an organ of protection. In: Fitzpatrick
TB, Eisen AZ, Wolff K, Freedberg IM, Austen KF, eds. Dermatology in
general medicine. 4th ed. Vol. 1. New York: McGraw-Hill, 1993:241-53.
2. Fearon DT, Locksley RM. The instructive role of innate immunity in
the acquired immune response. Science 1996;272:50-3.
3. Lehrer RI, Lichtenstein AK, Ganz T. Defensins: antimicrobial and cytotoxic peptides of mammalian cells. Annu Rev Immunol 1993;11:105-28.
4. Harder J, Bartels J, Christophers E, Schroder JM. A peptide antibiotic
from human skin. Nature 1997;387:861.
5. Stolzenberg ED, Anderson GM, Ackermann MR, Whitlock RH,
Zasloff M. Epithelial antibiotic induced in states of disease. Proc Natl Acad
Sci U S A 1997;94:8686-90.
6. Gallo RL, Ono M, Povsic T, et al. Syndecans, cell surface heparan sulfate proteoglycans, are induced by a proline-rich antimicrobial peptide from
wounds. Proc Natl Acad Sci U S A 1994;91:11035-9.
7. Frohm M, Agerberth B, Ahangari G, et al. The expression of the gene
coding for the antibacterial peptide LL-37 is induced in human keratinocytes during inflammatory disorders. J Biol Chem 1997;272:15258-63.
8. Fulton C, Anderson GM, Zasloff M, Bull R, Quinn AG. Expression of
natural peptide antibiotics in human skin. Lancet 1997;350:1750-1.
9. Gropp R, Frye M, Wagner TO, Bargon J. Epithelial defensins impair adenoviral infection: implication for adenovirus-mediated gene therapy. Hum
Gene Ther 1999;10:957-64.
10. Gallo RL, Huttner KM. Antimicrobial peptides: an emerging concept
in cutaneous biology. J Invest Dermatol 1998;111:739-43.
11. Nizet V, Ohtake T, Lauth X, et al. Innate antimicrobial peptide protects the skin from invasive bacterial infection. Nature 2001;414:454-7.
12. Panyutich A, Shi J, Boutz PL, Zhao C, Ganz T. Porcine polymorphonuclear leukocytes generate extracellular microbicidal activity by elastasemediated activation of secreted proprotegrins. Infect Immun 1997;65:97885.
13. Leung DYM. Atopic dermatitis: new insights and opportunities for
therapeutic intervention. J Allergy Clin Immunol 2000;105:860-76.
14. Leung DYM, Tharp M, Boguniewicz M. Atopic dermatitis (atopic eczema). In: Freedberg IM, Eisen AZ, Wolff K, et al., eds. Fitzpatricks dermatology in general medicine. 5th ed. Vol. 1. New York: McGraw-Hill,
1998:1464-80.
15. Christophers E, Henseler T. Contrasting disease patterns in psoriasis
and atopic dermatitis. Arch Dermatol Res 1987;279:Suppl:S48-S51.
16. Grice K, Sattar H, Baker H, Sharratt M. The relationship of transepidermal water loss to skin temperature in psoriasis and eczema. J Invest Dermatol 1975;64:313-5.
17. Dorschner RA, Pestonjamasp VK, Tamakuwala S, et al. Cutaneous injury induces the release of cathelicidin anti-microbial peptides active against
group A Streptococcus. J Invest Dermatol 2001;117:91-7.
18. Liu AY, Destoumieux D, Wong AV, et al. Human b-defensin-2 production in keratinocytes is regulated by interleukin-1, bacteria, and the
state of differentiation. J Invest Dermatol 2002;118:275-81.
N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org 1159
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
The Ne w E n g l a nd Jo u r n a l o f Me d ic i ne
19. Conover WJ. Practical nonparametric statistics. 2nd ed. New York:
John Wiley, 1980.
20. Peschel A, Jack RW, Otto M, et al. Staphylococcus aureus resistance to
human defensins and evasion of neutrophil killing via the novel virulence
factor MprF is based on modification of membrane lipids with L-lysine.
J Exp Med 2001;193:1067-76.
21. Leyden JJ, Kligman AM. The case for steroid-antibiotic combinations.
Br J Dermatol 1977;96:179-87.
22. Bunikowski R, Mielke ME, Skarabis H, et al. Evidence for a diseasepromoting effect of Staphylococcus aureus-derived exotoxins in atopic dermatitis. J Allergy Clin Immunol 2000;105:814-9.
23. Cho SH, Strickland I, Tomkinson A, Fehringer AP, Gelfand EW, Leung
DY. Preferential binding of Staphylococcus aureus to skin sites of Th2-mediated inflammation in a murine model. J Invest Dermatol 2001;116:658-63.
24. Nilsson EJ, Henning CG, Magnusson J. Topical corticosteroids and
Staphylococcus aureus in atopic dermatitis. J Am Acad Dermatol 1992;27:
29-34.
25. Remitz A, Kyllonen H, Granlund H, Reitamo S. Tacrolimus ointment
reduces staphylococcal colonization of atopic dermatitis lesions. J Allergy
Clin Immunol 2001;107:196-7.
26. Cho SH, Strickland I, Boguniewicz M, Leung DY. Fibronectin and
fibrinogen contribute to the enhanced binding of Staphylococcus aureus to
atopic skin. J Allergy Clin Immunol 2001;108:269-74.
27. Morishita Y, Tada J, Sato A, et al. Possible influences of Staphylococcus
aureus on atopic dermatitis the colonizing features and the effects of
staphylococcal enterotoxins. Clin Exp Allergy 1999;29:1110-7.
28. Giustizieri ML, Mascia F, Frezzolini A, et al. Keratinocytes from patients with atopic dermatitis and psoriasis show a distinct chemokine production profile in response to T cell-derived cytokines. J Allergy Clin Immunol 2001;107:871-7.
Copyright 2002 Massachusetts Medical Society.
1160 N Engl J Med, Vol. 347, No. 15 October 10, 2002 www.nejm.org
The New England Journal of Medicine
Downloaded from nejm.org on August 25, 2014. For personal use only. No other uses without permission.
Copyright 2002 Massachusetts Medical Society. All rights reserved.
Вам также может понравиться
- Hayashi Reiki ManualДокумент14 страницHayashi Reiki Manualboomerb100% (4)
- Osce Recalls 2011, 2013, 2016 2Документ6 страницOsce Recalls 2011, 2013, 2016 2FA Khan100% (1)
- Lotus BirthДокумент4 страницыLotus BirthindahОценок пока нет
- Chemotherapy Protocols V 101Документ42 страницыChemotherapy Protocols V 101Charm TanyaОценок пока нет
- IV FluidsДокумент3 страницыIV FluidsEvangeline San Jose83% (6)
- Vol 22.5 Neuroimaging.2016Документ460 страницVol 22.5 Neuroimaging.2016mikaaa000Оценок пока нет
- Basics of Microvascular SurgeryДокумент33 страницыBasics of Microvascular SurgeryPratikshya KothiaОценок пока нет
- Joost Schalkwijk., 2010 PDFДокумент11 страницJoost Schalkwijk., 2010 PDFInna LiashenkoОценок пока нет
- Immune Response Profiles in Human Skin: Epithelial DefenseДокумент7 страницImmune Response Profiles in Human Skin: Epithelial Defensematrixx15Оценок пока нет
- Allergic Skin DiseasesДокумент12 страницAllergic Skin DiseasesNo NameeОценок пока нет
- Atopic Dermatitis: From Pathophysiology To Diagnostic ApproachДокумент9 страницAtopic Dermatitis: From Pathophysiology To Diagnostic Approachwe sagara dewiОценок пока нет
- 440 SIG-1459 y SIG-1460: Nuevos Compuestos de Isoprenilcisteína AntiacnéДокумент1 страница440 SIG-1459 y SIG-1460: Nuevos Compuestos de Isoprenilcisteína AntiacnéFelipe Arango SotoОценок пока нет
- 90-Article Text With Names and Affiliation-154-1!10!20190919Документ4 страницы90-Article Text With Names and Affiliation-154-1!10!20190919Agharid Ali HusseinОценок пока нет
- TYPE IV HypersensitivityДокумент27 страницTYPE IV HypersensitivitylalaineОценок пока нет
- Kuliah ImunologiДокумент59 страницKuliah ImunologiUmmi Rinandari100% (1)
- Thermo-Responsive Triple-Function Nanotransporter For Efficient Chemo-Photothermal Therapy of Multidrug-Resistant Bacterial InfectionДокумент7 страницThermo-Responsive Triple-Function Nanotransporter For Efficient Chemo-Photothermal Therapy of Multidrug-Resistant Bacterial InfectiontasniaОценок пока нет
- Atopic Dermatitis .Документ7 страницAtopic Dermatitis .nyuwwchocolavaОценок пока нет
- Jin 2009 Models of Atopic DermatitisДокумент10 страницJin 2009 Models of Atopic DermatitisBrigitta SzöllősiОценок пока нет
- Immunological Mechanisms in Allergic Contact Dermatitis: Abstract and IntroductionДокумент11 страницImmunological Mechanisms in Allergic Contact Dermatitis: Abstract and IntroductionMarcellRaymondОценок пока нет
- International Immunopharmacology: SciencedirectДокумент11 страницInternational Immunopharmacology: SciencedirectHoàngОценок пока нет
- Iterleukin-4 and Interferon-Y in Allergic Contact Dermatitis With Atopic Background in Leather Tannery Factory WorkerДокумент6 страницIterleukin-4 and Interferon-Y in Allergic Contact Dermatitis With Atopic Background in Leather Tannery Factory WorkershaviraОценок пока нет
- Bucharest Vitamin D Cutolo PDFДокумент30 страницBucharest Vitamin D Cutolo PDFgeorgesculigiaОценок пока нет
- Skin Immunity To Dermatophytes: From Experimental Infection Models To Human DiseaseДокумент16 страницSkin Immunity To Dermatophytes: From Experimental Infection Models To Human Diseaseirwan junОценок пока нет
- Research Letters: Urticaria Caused by DimenhydrinateДокумент2 страницыResearch Letters: Urticaria Caused by DimenhydrinateNauroh NaurohОценок пока нет
- 2003 - Journal of Endodontics - 29 - 4 - 240 - 243 - Induction of Cyclooxygenase 2 mRNA and Protein Expressio PDFДокумент4 страницы2003 - Journal of Endodontics - 29 - 4 - 240 - 243 - Induction of Cyclooxygenase 2 mRNA and Protein Expressio PDFNamrata SachdevaОценок пока нет
- Aair 13 762Документ14 страницAair 13 762HELSINKI Alya FiskaОценок пока нет
- Lentiviral Vector-Mediated Correction of A Mouse Model of Leukocyte Adhesion Deficiency Type IДокумент11 страницLentiviral Vector-Mediated Correction of A Mouse Model of Leukocyte Adhesion Deficiency Type IDeede DarkoОценок пока нет
- Sumber Referat MataДокумент16 страницSumber Referat MatabethanurviaОценок пока нет
- Kimber 2002 PDFДокумент11 страницKimber 2002 PDFngekОценок пока нет
- Anetodermic Lupus Panniculitis and Antiphospholipid Antibodies: Report of Three CasesДокумент4 страницыAnetodermic Lupus Panniculitis and Antiphospholipid Antibodies: Report of Three CasesDermatologie VenerologieОценок пока нет
- Effects of Acupuncture On 1 Chloro 2 4 Dinitrochlo - 2017 - Journal of Acupunctu PDFДокумент9 страницEffects of Acupuncture On 1 Chloro 2 4 Dinitrochlo - 2017 - Journal of Acupunctu PDFAugustusОценок пока нет
- Innate Immunity, Microbiology, Inflammation - ABSTRACTSДокумент1 страницаInnate Immunity, Microbiology, Inflammation - ABSTRACTSFebrian AlfaroОценок пока нет
- Gao 2012Документ8 страницGao 2012GHUJОценок пока нет
- Infect. Immun.-2002-Bacellar-6734-40Документ7 страницInfect. Immun.-2002-Bacellar-6734-40Pedro Henrique De MirandaОценок пока нет
- Skin Microbiome in Atopic DermatitisДокумент9 страницSkin Microbiome in Atopic Dermatitisdr_miguelОценок пока нет
- Effector and Regulatory Mechanisms in Allergic Contact DermatitisДокумент16 страницEffector and Regulatory Mechanisms in Allergic Contact Dermatitistri erdiansyahОценок пока нет
- Development of An Enzyme-Linked Immunosorbant Assay (ELISA) For The Serodiagnosis of Canine Dermatophytosis Caused by Microsporum CanДокумент6 страницDevelopment of An Enzyme-Linked Immunosorbant Assay (ELISA) For The Serodiagnosis of Canine Dermatophytosis Caused by Microsporum CanjenОценок пока нет
- Chapter 13:: Basic Principles of Immunologic Diseases in Skin (Pathophysiology of Immunologic/Inflammatory Skin Diseases)Документ7 страницChapter 13:: Basic Principles of Immunologic Diseases in Skin (Pathophysiology of Immunologic/Inflammatory Skin Diseases)S FznsОценок пока нет
- Out 3Документ9 страницOut 3Musthafa Afif WardhanaОценок пока нет
- Nihms891003. MicrobiomaДокумент22 страницыNihms891003. MicrobiomaEugeniaОценок пока нет
- Evaluation of Itch by Using Nc/Ngatnd Mice: A Model of Human Atopic DermatitisДокумент8 страницEvaluation of Itch by Using Nc/Ngatnd Mice: A Model of Human Atopic DermatitisDewiAsfarОценок пока нет
- Interleukin-17 IL-17-mediated Immunity Controls SKДокумент11 страницInterleukin-17 IL-17-mediated Immunity Controls SKnadyasantikaОценок пока нет
- Research Article Cell Response: β-Defensin Strengthens Antimicrobial Peritoneal MastДокумент14 страницResearch Article Cell Response: β-Defensin Strengthens Antimicrobial Peritoneal MastVictor Hugo DelgadilloОценок пока нет
- Biomedicine - Atopy 2021 0115 1355Документ18 страницBiomedicine - Atopy 2021 0115 1355Quynh NguyenОценок пока нет
- Autoantibodies Against The Processed Ectodomain of Collagen Xvii (Bpag2, Bp180) Define A Canine Homologue of Linear Iga Disease of HumansДокумент8 страницAutoantibodies Against The Processed Ectodomain of Collagen Xvii (Bpag2, Bp180) Define A Canine Homologue of Linear Iga Disease of HumansLauОценок пока нет
- Melendres Chem-103 Final-Requirements May-2022Документ31 страницаMelendres Chem-103 Final-Requirements May-2022Bianca MelendresОценок пока нет
- Expressions of Antimicrobial Peptides LL 37, Human Beta Defensin 2 and 3 in The Lesions of Cutaneous Tuberculosis and TuberculidsДокумент6 страницExpressions of Antimicrobial Peptides LL 37, Human Beta Defensin 2 and 3 in The Lesions of Cutaneous Tuberculosis and TuberculidsRiefka Ananda ZulfaОценок пока нет
- Cryptococcal Cellulitis in A Patient With Myelodysplastic SyndromeДокумент2 страницыCryptococcal Cellulitis in A Patient With Myelodysplastic SyndromeSan Maria SitompulОценок пока нет
- Tosca 1996Документ7 страницTosca 1996HEARTJUGGLERОценок пока нет
- Specific Immunotherapy in Atopic Dermatitis: ReviewДокумент9 страницSpecific Immunotherapy in Atopic Dermatitis: ReviewEsty GusmelisaОценок пока нет
- Dermatology: World Journal ofДокумент6 страницDermatology: World Journal ofRika FitriaОценок пока нет
- Cytokine and Growth Factor Reviews: SciencedirectДокумент10 страницCytokine and Growth Factor Reviews: SciencedirectluciaОценок пока нет
- Article 1- מחקר עורДокумент15 страницArticle 1- מחקר עורnoam tiroshОценок пока нет
- Art 3A10.1186 2F1476 9255 5 16 - 2Документ11 страницArt 3A10.1186 2F1476 9255 5 16 - 2Lilis RohaetiОценок пока нет
- Jurnal HibridisasiДокумент12 страницJurnal HibridisasiNurfanida Natasya mОценок пока нет
- Novak Et Al 2011 AllergyДокумент10 страницNovak Et Al 2011 AllergyArturo VeraОценок пока нет
- BMC Dermatology: Association of Toll-Interacting Protein Gene Polymorphisms With Atopic DermatitisДокумент8 страницBMC Dermatology: Association of Toll-Interacting Protein Gene Polymorphisms With Atopic DermatitisdermRounds Dermatology NetworkОценок пока нет
- Techniques FOR Evaluating Humoral AND Cell-Mediated Immunity IN Mule Deer FawnsДокумент7 страницTechniques FOR Evaluating Humoral AND Cell-Mediated Immunity IN Mule Deer Fawnsrocky00023Оценок пока нет
- Eczema, Immunity & The Skin Microbiome: Heidi H. Kong Dermatology Branch, CCR, NCIДокумент38 страницEczema, Immunity & The Skin Microbiome: Heidi H. Kong Dermatology Branch, CCR, NCIrafdiaz2Оценок пока нет
- Patch Testing of Experimentally Sensitized Beagle Dogs - Development of A Model For Skin Lesions of Atopic Dermatitis (Pages 95-102)Документ8 страницPatch Testing of Experimentally Sensitized Beagle Dogs - Development of A Model For Skin Lesions of Atopic Dermatitis (Pages 95-102)jenОценок пока нет
- Jced 7 E656Документ4 страницыJced 7 E656María José VázquezОценок пока нет
- 2020 - Pemphigus Vulgaris and Bullous Pemphigoid Update On Diagnosis and TreatmentДокумент12 страниц2020 - Pemphigus Vulgaris and Bullous Pemphigoid Update On Diagnosis and TreatmentnancyerlenОценок пока нет
- 05.immune Mechanisms Leading To AtopicДокумент12 страниц05.immune Mechanisms Leading To AtopicVARUN BHARADWAJ .MОценок пока нет
- 810 jmm000095Документ8 страниц810 jmm000095Aditya Yudha PratamaОценок пока нет
- Molecular Pathogenesis of LeprosyДокумент5 страницMolecular Pathogenesis of LeprosyyournightОценок пока нет
- In Vitro Study of Photodynamic Therapy For Treatment of Bacteremia in Whole BloodДокумент7 страницIn Vitro Study of Photodynamic Therapy For Treatment of Bacteremia in Whole BloodSabrina JonesОценок пока нет
- MeniereДокумент8 страницMeniereTaqwa NindaОценок пока нет
- Skin and Soft-Tissue Infections CausedДокумент11 страницSkin and Soft-Tissue Infections CausedAni IoanaОценок пока нет
- Early MarriageДокумент9 страницEarly MarriageTaqwa NindaОценок пока нет
- AДокумент9 страницATaqwa NindaОценок пока нет
- Pediatrics 2008 Krakowski 812 24Документ15 страницPediatrics 2008 Krakowski 812 24Taqwa NindaОценок пока нет
- Flaherty 2012 CombinedДокумент10 страницFlaherty 2012 CombinedTaqwa NindaОценок пока нет
- Endogenous Antimicrobial Peptides and Skin Infections in Atopic DermatitisДокумент10 страницEndogenous Antimicrobial Peptides and Skin Infections in Atopic DermatitisTaqwa NindaОценок пока нет
- Jurnal Bells PalsyДокумент10 страницJurnal Bells PalsyputriОценок пока нет
- JaundiceДокумент19 страницJaundiceketsela12Оценок пока нет
- 2009 11 Oral GalvanismДокумент2 страницы2009 11 Oral GalvanismSppatilОценок пока нет
- CombinedДокумент1 страницаCombinedgldiatrОценок пока нет
- Curriculum Vitae Dr. Ari AstramДокумент4 страницыCurriculum Vitae Dr. Ari AstramBedah ManadoОценок пока нет
- Basic and Clinical Pharmacology 12th Edition-Bertram Katzung Susan Masters Anthony Trevor-290-296 PDFДокумент7 страницBasic and Clinical Pharmacology 12th Edition-Bertram Katzung Susan Masters Anthony Trevor-290-296 PDFalinamatei1000000Оценок пока нет
- Respiratory PhysiologyДокумент9 страницRespiratory PhysiologyBrent TorresОценок пока нет
- TrypanosomiasisДокумент39 страницTrypanosomiasisNani HendrianiОценок пока нет
- MHRA Warning Letter Metal Metal 1Документ4 страницыMHRA Warning Letter Metal Metal 1Ron WoeringОценок пока нет
- Microsurgical Training With FreshДокумент5 страницMicrosurgical Training With FreshBob sponjaОценок пока нет
- PEH 3 Lesson 2Документ3 страницыPEH 3 Lesson 2ShaineMaiko MarigocioОценок пока нет
- Parsial Hidrolisat PikabДокумент25 страницParsial Hidrolisat PikabJulistio DjaisОценок пока нет
- Clinical Session Adult I-1 3Документ34 страницыClinical Session Adult I-1 3Juan LinОценок пока нет
- MCN 2Документ106 страницMCN 2Nicole Dela TorreОценок пока нет
- Root BiomodificationДокумент88 страницRoot Biomodificationperiodontics07Оценок пока нет
- Literature Review 1Документ5 страницLiterature Review 1api-550490262Оценок пока нет
- Prudential BSN Takaful Berhad Investment-Linked Plan IllustrationДокумент19 страницPrudential BSN Takaful Berhad Investment-Linked Plan IllustrationhilmiyaidinОценок пока нет
- Immunization Schedule in India 2017 (Latest !!)Документ13 страницImmunization Schedule in India 2017 (Latest !!)rajОценок пока нет
- GAD Aos 40 AnosДокумент7 страницGAD Aos 40 AnosrafaelplОценок пока нет
- Drug StudyДокумент5 страницDrug Studyanon_168410816Оценок пока нет
- TraumaAirway Management - ValДокумент36 страницTraumaAirway Management - ValValerie Suge-MichiekaОценок пока нет
- Pengaruh Inisiasi Menyusu Dini Terhadap Jumlah Perdarahan Pasca PersalinanДокумент10 страницPengaruh Inisiasi Menyusu Dini Terhadap Jumlah Perdarahan Pasca Persalinanstasebolen asikОценок пока нет
- Quarter 3 Study Guide: Digestive SystemДокумент3 страницыQuarter 3 Study Guide: Digestive SystemChristina TangОценок пока нет