Академический Документы
Профессиональный Документы
Культура Документы
EPICH 16 Masami Computer PC
Загружено:
Roberto Alfredo SáenzАвторское право
Доступные форматы
Поделиться этим документом
Поделиться или встроить документ
Этот документ был вам полезен?
Это неприемлемый материал?
Пожаловаться на этот документАвторское право:
Доступные форматы
EPICH 16 Masami Computer PC
Загружено:
Roberto Alfredo SáenzАвторское право:
Доступные форматы
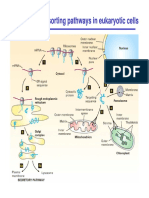
Three dimensional promoter interactions
during a circadian cycle
Ando-Kuri M1, Morf Jorg2, Wingett Steven2, Crains Jonathan2, Fraser Peter2, Furlan-Magaril Mayra1,2.
1 Instituto de Fisiología Celular, Universidad Nacional Autónoma de México, Mexico City, Mexico.
Abstract 2 Nuclear Dynamics Programme, Babraham Institute, Cambridge, CB22 3AT UK
To have a better understanding of how chromosome
conformation influences transcriptional regulation it is useful Promoter interacting regions are enriched in Phase correlation between circadian promoter-
to analyze 3D chromatin organization in transcriptionally enhancer features including eRNAs eRNAs interacting pairs
dynamic situations in a postmitotic tissue such as a circadian
*** 22-24
cycle in mouse liver. We applied Capture-HiC (CHiC) to study R2=0.06165394
9-21
chromosome conformation between all gene promoters and Observed
their associated genomic elements providing a map of Expected 6-18
significant interactions at 4 time points during the course of a 3-15
Phases
circadian cycle. ***
Interactions
0-12
We found that genomic regions interacting with ***
promoters of circadian genes not only show enhancer-specific 7-9
eRNAs
histone modifications but often exhibit oscillatory enhancer *** 4-6 Circadian
RNA expression. Interestingly, the peak expression for most *** promoters
1-3
eRNAs is timed around dawn while gene expression of
contacted circadian genes show an even phase distribution Pairs
around the clock. Furthermore, in general we detect no Fig. 5. Phase distribution of promoter (purple) and eRNA (orange)
eRNAs H3K27ac H3K4me1 H3K4me3 CTCF
temporal variation in interaction frequency between circadian interacting pairs. Each column contains one pair.
Fig. 2. Enrichment of histone modifications, CTCF and eRNAs at
promoters and other genomic regions. This suggests a model promoter interacting regions.
in which topological scaffolds install regulatory contacts for
circadian gene expression but other factors than spatial
chromatin organization and eRNAs determine circadian Circadian enhancers Core clock gene Interaction profiles from circadian gene
phase of transcription. preferentially contact promoters form less promoters remain mostly stable over time
circadian gene interactions than non
Experimental Strategy: Promoter promoters core clock genes. Npas2
a b
Capture-HiC *** Observed
Expected
Non core-clock genes
Core clock genes
eRNAs
ZT0
Circadian promoters
CHiC is a novel method to enrich interactions of interest from
a HiC library,1,2 (Fig. 1). Promoter-CHiC was performed in Interactions
*
triplicate at ZT0, 6, 12 and 18 during a circadian cycle in
mouse adult liver. Circadian gene expression was confirmed ZT6
by RNAseq on the same samples (not shown).
Hi-C library
Oscillatory eRNAs CHiC interactions
Promoter B
Fig. 3. a) The expected number interactions was estimated by ZT12
Promoter generating a distribution for random oscillatory eRNAs b) Average
number of interactions from 25 core clock genes.
Sequence capture Streptavidin pulldown
hybridization
Circadian promoters interact with oscillatory ZT18
B
B
Promoter B enhancers with peak expression around dawn
B RNA probe
Promoter a
B
Associated circadian
PCR Fig. 6. The Npas2 gene promoter interaction landscape. Npas2 promoter
ZT0 contacts a circadian eRNA enhancer and the interaction is stable over
C-HiC Library
promoters
time.
Bait OtherEnd
ZT18 ZT6
Conclusions
ZT12 ● Enhancer features and eRNAs are enriched at the significant
Fig. 1. Capture-HiC protocol. RNA biotinylated baits are used to pull down interactions detected by promoter CHiC.
all ligation products containing a promoter on either side of the interaction. b eRNAs phases
● Core clock gene promoters make significantly less contacts
than other oscillatory genes which suggests their
Results transcriptional regulation might not require distal elements.
● Circadian gene promoters in all transcriptional phases
Significant interactions were detected using CHiCAGO preferentially interact with oscillating eRNAs expressed in
pipeline (Capture HiC Analysis of Genomic ZT18-ZT3.
Organization)3. CHiC retrieved ~150,000 interactions per ● Overall, the interaction profiles from circadian gene promoters
timepoint with ~10 interactions on average per gene are maintained over time providing a spacial framework that
allows efficient circadian gene expression control
promoter.
We found an enrichment of enhancer features over the
Diurnal promoters Nocturnal promoters
1Schoenfelder, S., Furlan-Magaril, M., Mifsud, B., Tavares-Cadete, F., Sugar, R., Javierre, B. M., ... & Dimitrova, E. (2015). The pluripotent regulatory circuitry connecting
promoter interacting regions (P < 0.001 t-test, Fig. 2) and promoters to their long-range interacting elements. Genome research, 25(4), 582-597.
2Mifsud, B., Tavares-Cadete, F., Young, A. N., Sugar, R., Schoenfelder, S., Ferreira, L., ... & Herman, B. (2015). Mapping long-range promoter contacts in human cells with
Fig. 4. a) Number of circadian promoters interacting with oscillating high-resolution capture Hi-C. Nature genetics, 47(6), 598-606.
occupancy of circadian transcription factors (P < 0.001 t-test, enhancers b) Phase distribution of eRNAs elements interacting with
3Cairns, J., Freire-Pritchett, P., Wingett, S. W., Dimond, A., Plagnol, V., Zerbino, D., ... & Spivakov, M. (2015). CHiCAGO: robust detection of DNA looping interactions in
capture Hi-C data. bioRxiv, 028068.
4Fang, B., Everett, L. J., Jager, J., Briggs, E., Armour, S. M., Feng, D., ... & Lazar, M. A. (2014). Circadian enhancers coordinate multiple phases of rhythmic gene
not shown). diurnal and nocturnal circadian promoters. transcription in vivo. Cell, 159(5), 1140-1152.
Вам также может понравиться
- 1 s2.0 S1874939919302160 MainДокумент11 страниц1 s2.0 S1874939919302160 MainxОценок пока нет
- Project BoardДокумент3 страницыProject BoardPariОценок пока нет
- Genetics: From Genes To GenomesДокумент58 страницGenetics: From Genes To GenomesAshleyОценок пока нет
- Cellular Functions of Long Noncoding Rnas: Run-Wen Yao, Yang Wang and Ling-Ling ChenДокумент10 страницCellular Functions of Long Noncoding Rnas: Run-Wen Yao, Yang Wang and Ling-Ling ChenAmrutha Swathi PriyaОценок пока нет
- Unbiased Interrogation of 3D Genome Topology Using Chromosome Conformation Capture Coupled To High-Throughput Sequencing (4C-Seq)Документ22 страницыUnbiased Interrogation of 3D Genome Topology Using Chromosome Conformation Capture Coupled To High-Throughput Sequencing (4C-Seq)Sophia OlveraОценок пока нет
- 2006-Slicer and The ArgonautesДокумент8 страниц2006-Slicer and The ArgonautesJorge Hantar Touma LazoОценок пока нет
- Nature 23 Complementary Alu Sequences Mediate Enhancer-Promoter SelectivityДокумент32 страницыNature 23 Complementary Alu Sequences Mediate Enhancer-Promoter Selectivitylandau1994Оценок пока нет
- Mine - SylBioGr10Документ11 страницMine - SylBioGr10Suzan.FilizОценок пока нет
- Protein SynthesisДокумент19 страницProtein SynthesisRara Aulia IIОценок пока нет
- 5 Systems and Synthetic microRNA BiologyДокумент23 страницы5 Systems and Synthetic microRNA BiologySandra GonzalezОценок пока нет
- Nearest-Neighbor E Ffects Modulate Loxp Spacer Dna Chemical Shifts and Guide Oligonucleotide Design For Nuclear Magnetic Resonance StudiesДокумент10 страницNearest-Neighbor E Ffects Modulate Loxp Spacer Dna Chemical Shifts and Guide Oligonucleotide Design For Nuclear Magnetic Resonance StudiesRenan Guilherme de Oliveira GuihОценок пока нет
- Syl Bio GR 10Документ5 страницSyl Bio GR 10Suzan FilizОценок пока нет
- Kimed 1Документ15 страницKimed 1fifin oktavianiОценок пока нет
- Interactions Between Short and Long Noncoding RnasДокумент10 страницInteractions Between Short and Long Noncoding RnasxОценок пока нет
- Li Et Al 2017 Magnesium Catalysis Mediated Tetrazoles in Desymmetrization Reaction of AziridinesДокумент4 страницыLi Et Al 2017 Magnesium Catalysis Mediated Tetrazoles in Desymmetrization Reaction of AziridinesNoimurОценок пока нет
- Topic 24Документ54 страницыTopic 24bpwalshОценок пока нет
- Research Article: Trends in The Binding of Cell Penetrating Peptides To Sirna: A Molecular Docking StudyДокумент13 страницResearch Article: Trends in The Binding of Cell Penetrating Peptides To Sirna: A Molecular Docking StudyalfaОценок пока нет
- 1 s2.0 S0968000415001644 MainДокумент11 страниц1 s2.0 S0968000415001644 MainTheresa LambayonОценок пока нет
- Exam 3 Study SheetДокумент4 страницыExam 3 Study Sheetapi-659209201Оценок пока нет
- Micrornas: Key Regulators of Stem Cells: Vamsi K. Gangaraju and Haifan LinДокумент10 страницMicrornas: Key Regulators of Stem Cells: Vamsi K. Gangaraju and Haifan LinEdgar Huerta CardenasОценок пока нет
- Encapsulation State of Messenger RNA Inside Lipid NanoparticlesДокумент5 страницEncapsulation State of Messenger RNA Inside Lipid NanoparticlesPencari IlmuОценок пока нет
- SylBioGr10 - Updated - FinalДокумент9 страницSylBioGr10 - Updated - FinalSuzan FilizОценок пока нет
- November 10Документ28 страницNovember 10Tara BhatnagarОценок пока нет
- Chapter 4Документ61 страницаChapter 4mlОценок пока нет
- Pre-mRNA Splicing Life at The Centre of The Central DogmaДокумент3 страницыPre-mRNA Splicing Life at The Centre of The Central DogmaQwyn Kym De GuzmanОценок пока нет
- Rab Proteins As Membrane Organizers: Marino Zerial and Heidi McbrideДокумент13 страницRab Proteins As Membrane Organizers: Marino Zerial and Heidi McbrideCourtney JohnsonОценок пока нет
- (Sample Open Access Content) : Methodology ArticleДокумент7 страниц(Sample Open Access Content) : Methodology ArticleMari SuganthiОценок пока нет
- Structure and Function of RNA - Microbiology - OpenStaxДокумент5 страницStructure and Function of RNA - Microbiology - OpenStaxAleksandra Sanja MartinovicОценок пока нет
- Régulation de L'éxpréssion Génétique (1) - CopieДокумент163 страницыRégulation de L'éxpréssion Génétique (1) - CopieWahuba RahmaniОценок пока нет
- Lec7 1pptДокумент21 страницаLec7 1pptShannon MarieОценок пока нет
- microRNAs-based Therapeutics in - Neurodegenerative DiseasesДокумент26 страницmicroRNAs-based Therapeutics in - Neurodegenerative DiseasesamyОценок пока нет
- Nuclear Receptors: G-Protein CoupledДокумент2 страницыNuclear Receptors: G-Protein CoupledJordan DouglasОценок пока нет
- Regulation of Nuclear RNA Exosome ComplexДокумент13 страницRegulation of Nuclear RNA Exosome ComplexElgarОценок пока нет
- Cell Biology Presentation: Group-4 Concept of Ribozyme and Rna TransductionДокумент38 страницCell Biology Presentation: Group-4 Concept of Ribozyme and Rna TransductionRafia AslamОценок пока нет
- Chen (2020)Документ15 страницChen (2020)IVAN LUIS FERNANDEZ DE LA CRUZОценок пока нет
- Harper S Illustrated Biochemistry by Vic-385-403Документ19 страницHarper S Illustrated Biochemistry by Vic-385-403DavidОценок пока нет
- Access: Biswajit Biswas and Prashant Chandra SinghДокумент9 страницAccess: Biswajit Biswas and Prashant Chandra SinghSourashis BiswasОценок пока нет
- Resource: Engineering Complex Synthetic Transcriptional Programs With CRISPR RNA ScaffoldsДокумент12 страницResource: Engineering Complex Synthetic Transcriptional Programs With CRISPR RNA ScaffoldsLetícia AlibertiОценок пока нет
- Youssef Et Al 2023 Informational Polymers With Precise Carbamate SequencesДокумент5 страницYoussef Et Al 2023 Informational Polymers With Precise Carbamate SequencesPascal Niño RodriguezОценок пока нет
- YEBEДокумент14 страницYEBELegeek GamingОценок пока нет
- RSC Advances: PaperДокумент14 страницRSC Advances: PaperDavid RincónОценок пока нет
- Jacs 0c11605Документ13 страницJacs 0c11605Mérito MéritoОценок пока нет
- The Target of Rapamycin Signalling Pathway in Ageing and Lifespan RegulationДокумент21 страницаThe Target of Rapamycin Signalling Pathway in Ageing and Lifespan Regulationmdjgyv1933Оценок пока нет
- Targeting Polycomb Systems To Regulate Gene ExpressionДокумент7 страницTargeting Polycomb Systems To Regulate Gene ExpressiondffОценок пока нет
- RSC Advances: PaperДокумент7 страницRSC Advances: PaperDiego Alejandro Hurtado BalcazarОценок пока нет
- Final Capstone Poster-Mann PDFДокумент1 страницаFinal Capstone Poster-Mann PDFAmy Nottingham-MartinОценок пока нет
- TMP 583 EДокумент6 страницTMP 583 EFrontiersОценок пока нет
- 5) Cell Structure Summary - 9744 - 2018Документ2 страницы5) Cell Structure Summary - 9744 - 2018GUCCINOОценок пока нет
- Org. Lett. 2019, 21, 10115-10119Документ5 страницOrg. Lett. 2019, 21, 10115-10119NoimurОценок пока нет
- F22 MCB 2050 Lecture 2 - Polymerases and PromotersДокумент22 страницыF22 MCB 2050 Lecture 2 - Polymerases and PromotersNO VIDEOSОценок пока нет
- 1 s2.0 S2666166722004853 MainДокумент33 страницы1 s2.0 S2666166722004853 Mainjiahuan.he90Оценок пока нет
- 2018 DestaracДокумент10 страниц2018 DestaracMarion ChenalОценок пока нет
- Walter Gilbert RNA WorldДокумент1 страницаWalter Gilbert RNA WorldARGHA MANNAОценок пока нет
- Unwinding Rna'S Secrets: Advances in The Biology, Physics, and Modeling of Complex RnasДокумент10 страницUnwinding Rna'S Secrets: Advances in The Biology, Physics, and Modeling of Complex Rnasa4agarwalОценок пока нет
- QSB 03 - Chemical Components and Energy2Документ30 страницQSB 03 - Chemical Components and Energy2fta2013Оценок пока нет
- Trna Model PDFДокумент3 страницыTrna Model PDFYoga NovalОценок пока нет
- A Hierarchical Model For Evolutions of 23S Ribosomal RNA - Konstantin Bokov - Sergey V Steinberg - Nature - 2011Документ4 страницыA Hierarchical Model For Evolutions of 23S Ribosomal RNA - Konstantin Bokov - Sergey V Steinberg - Nature - 2011Antonio Vázquez MotaОценок пока нет
- 4-BIOL 101 Study Guide Quiz 4Документ5 страниц4-BIOL 101 Study Guide Quiz 4Suraj NaikОценок пока нет
- Circ InteractomeДокумент9 страницCirc InteractomeBahlibiОценок пока нет
- Radar Target Detection: Handbook of Theory and PracticeОт EverandRadar Target Detection: Handbook of Theory and PracticeРейтинг: 5 из 5 звезд5/5 (1)
- Problemas de Limites, Continuidad y DerivabilidadДокумент42 страницыProblemas de Limites, Continuidad y DerivabilidadCecilia Alvarez HerreraОценок пока нет
- Cancers 11 00061Документ18 страницCancers 11 00061Roberto Alfredo SáenzОценок пока нет
- Nutrient Value BookletДокумент68 страницNutrient Value Bookletsirius8Оценок пока нет
- Schedule 5 Trial +15 WeeksДокумент3 страницыSchedule 5 Trial +15 WeeksAugusto MigliorinОценок пока нет
- Genomic Interactome of Circadian Genes in Mouse TissuesДокумент1 страницаGenomic Interactome of Circadian Genes in Mouse TissuesRoberto Alfredo SáenzОценок пока нет
- Greyhound - Ca - Step 4Документ3 страницыGreyhound - Ca - Step 4Roberto Alfredo SáenzОценок пока нет
- Plate Grid Setup Report: Experiment: 20170315 - Bmal1 - Rps9 - Standard - Curv Applied BiosystemsДокумент1 страницаPlate Grid Setup Report: Experiment: 20170315 - Bmal1 - Rps9 - Standard - Curv Applied BiosystemsRoberto Alfredo SáenzОценок пока нет
- RNA Seq TutorialДокумент139 страницRNA Seq TutorialRoberto Alfredo Sáenz0% (1)
- Minorities and IndegeniousДокумент262 страницыMinorities and IndegeniousRoberto Alfredo SáenzОценок пока нет
- What Is To Be Done PDFДокумент488 страницWhat Is To Be Done PDFfluxosОценок пока нет
- Extraction - Lab Fall 2010aДокумент6 страницExtraction - Lab Fall 2010aRoberto Alfredo SáenzОценок пока нет
- Why We Are Marxists - in Defence of MarxismДокумент6 страницWhy We Are Marxists - in Defence of MarxismRoberto Alfredo SáenzОценок пока нет
- USPI - Bicillin C-R - Penicillin G Benzathine, Penicillin GДокумент11 страницUSPI - Bicillin C-R - Penicillin G Benzathine, Penicillin GRoberto Alfredo SáenzОценок пока нет
- Vibrational Assignments Thio-Tetrazole JMS03Документ11 страницVibrational Assignments Thio-Tetrazole JMS03Roberto Alfredo SáenzОценок пока нет
- Formulation and Evaluation of Herbal Lip Rouge.: Research ArticleДокумент5 страницFormulation and Evaluation of Herbal Lip Rouge.: Research ArticleTynОценок пока нет
- Daftar PustakaДокумент5 страницDaftar PustakaAnonymous ASaqa0E0qgОценок пока нет
- Acid Base PhysiologyДокумент1 страницаAcid Base PhysiologyHAMMYER ALROKHAMIОценок пока нет
- LAN Party Skate Park by Shane Jesse ChristmassДокумент91 страницаLAN Party Skate Park by Shane Jesse ChristmassPatrick TrottiОценок пока нет
- The Retention of Complete DenturesДокумент12 страницThe Retention of Complete DentureswindOwhispersОценок пока нет
- Beatriz Colomina X Ray Architecture Lars Muller (046 090)Документ45 страницBeatriz Colomina X Ray Architecture Lars Muller (046 090)SABUESO FINANCIEROОценок пока нет
- Carbon Monoxide PoisoningДокумент31 страницаCarbon Monoxide PoisoningDheerajОценок пока нет
- Bedwetting in ChildrenДокумент41 страницаBedwetting in ChildrenAnkit ManglaОценок пока нет
- Why Jaggery? Is Jaggery Healthy? What Is Better: Jaggery or Sugar? Can Jaggery Cure Ailments?Документ5 страницWhy Jaggery? Is Jaggery Healthy? What Is Better: Jaggery or Sugar? Can Jaggery Cure Ailments?satheb319429Оценок пока нет
- DR Ololade 038Документ13 страницDR Ololade 038Andrawus DanjumaОценок пока нет
- Cast and SplintsДокумент58 страницCast and SplintsSulabh Shrestha100% (2)
- NURSING CARE PLAN - Decreased Cardiac OutputДокумент2 страницыNURSING CARE PLAN - Decreased Cardiac OutputRy Pablo83% (41)
- A Manual 20199 CompleetДокумент201 страницаA Manual 20199 CompleetJanPerezОценок пока нет
- PowersДокумент14 страницPowersIvan SokolovОценок пока нет
- ZFN, TALEN, and CRISPR-Cas-based Methods For Genome EngineeringДокумент9 страницZFN, TALEN, and CRISPR-Cas-based Methods For Genome EngineeringRomina Tamara Gil RamirezОценок пока нет
- Nur Writing - Marilyn JohnsonДокумент4 страницыNur Writing - Marilyn Johnsonyinghua guo0% (1)
- ERCPДокумент4 страницыERCPbilly wilsonОценок пока нет
- Slcog Abstracts 2012Документ133 страницыSlcog Abstracts 2012Indika WithanageОценок пока нет
- Cholecystitis BelgradeДокумент52 страницыCholecystitis BelgradeLazar VučetićОценок пока нет
- Polydactyly of The Foot A Review.92Документ10 страницPolydactyly of The Foot A Review.92mamyeu1801Оценок пока нет
- 2018 Surgical Rescue in Medical PatientsДокумент11 страниц2018 Surgical Rescue in Medical PatientsgiseladlrОценок пока нет
- Advanced Handbook of Systemic Lupus ErythematosusДокумент179 страницAdvanced Handbook of Systemic Lupus ErythematosusCésar CuadraОценок пока нет
- Operator MGC Ultima PFX - English - 142152-001rCДокумент65 страницOperator MGC Ultima PFX - English - 142152-001rCEvangelosОценок пока нет
- Zoology MCQs Practice Test 19Документ4 страницыZoology MCQs Practice Test 19Rehan HaiderОценок пока нет
- COPD CurrentДокумент9 страницCOPD Currentmartha kurniaОценок пока нет
- IMTX PatentДокумент76 страницIMTX PatentCharles GrossОценок пока нет
- Biology Combined Science Notes: Changamire Farirayi D.T. Form 3 & 4 BiologyДокумент67 страницBiology Combined Science Notes: Changamire Farirayi D.T. Form 3 & 4 BiologyKeiron Cyrus100% (6)
- Management of Imature ApexДокумент21 страницаManagement of Imature ApexBogdanОценок пока нет
- Expression of Interest - Elective Placement For Medical StudentsДокумент2 страницыExpression of Interest - Elective Placement For Medical Studentsfsdfs100% (1)
- FOCGB3 Utest DLR 6BДокумент2 страницыFOCGB3 Utest DLR 6BGyurácz GyulaОценок пока нет